ChIP-seq vs EMSA: Choosing the Right Tool for Transcription Factor Binding Analysis in Research & Drug Development
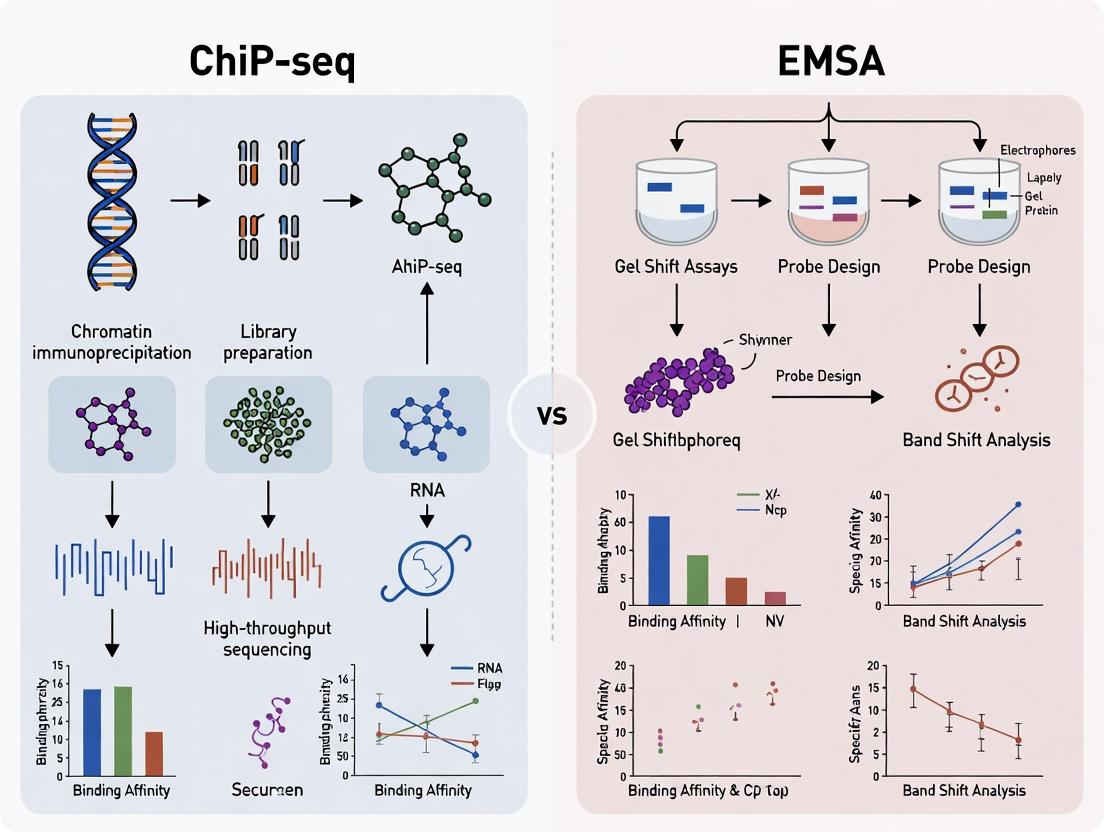
This comprehensive guide compares Chromatin Immunoprecipitation Sequencing (ChIP-seq) and Electrophoretic Mobility Shift Assay (EMSA), two pivotal techniques for studying transcription factor (TF)-DNA interactions.
ChIP-seq vs EMSA: Choosing the Right Tool for Transcription Factor Binding Analysis in Research & Drug Development
Abstract
This comprehensive guide compares Chromatin Immunoprecipitation Sequencing (ChIP-seq) and Electrophoretic Mobility Shift Assay (EMSA), two pivotal techniques for studying transcription factor (TF)-DNA interactions. Tailored for researchers, scientists, and drug development professionals, the article provides a foundational understanding of each method, explores their specific methodological applications and workflow requirements, addresses common troubleshooting and optimization challenges, and offers a direct, data-driven comparison for validation strategies. By synthesizing current best practices and limitations, this article serves as a strategic resource for selecting and implementing the optimal approach to elucidate gene regulatory mechanisms in basic research and therapeutic target discovery.
Understanding the Basics: What Are ChIP-seq and EMSA and How Do They Work?
The study of transcription factor (TF)-DNA interactions is fundamental to understanding gene regulation. Two primary techniques dominate this field: Chromatin Immunoprecipitation followed by sequencing (ChIP-seq) and the Electrophoretic Mobility Shift Assay (EMSA). While ChIP-seq identifies TF binding sites across the genome in vivo, EMSA provides a complementary, in vitro approach to probe the direct, biophysical interactions between a purified protein and a specific DNA sequence. This whitepaper details the core principles, protocols, and applications of EMSA, framing it as an essential tool for validating and mechanistically dissecting interactions suggested by high-throughput in vivo methods like ChIP-seq.
Core Principle of EMSA
The Electrophoretic Mobility Shift Assay (EMSA), also known as a gel shift assay, is based on a simple principle: the formation of a protein-nucleic acid complex reduces its mobility during non-denaturing polyacrylamide or agarose gel electrophoresis compared to the free nucleic acid probe. This "shift" in migration is detectable via autoradiography, fluorescence, or chemiluminescence.
Key Interactions Probed:
- Protein-DNA: Direct binding of TFs, nucleosomes, or other DNA-binding proteins.
- Protein-RNA: Binding of RNA-binding proteins.
- Complex Assembly: Formation of higher-order nucleoprotein complexes.
Primary Advantages over ChIP-seq:
- Direct Interaction Proof: Confirms a protein binds DNA directly, without requiring cellular machinery or antibody specificity.
- Quantitative Binding Parameters: Can determine dissociation constants (Kd) and binding stoichiometry.
- Mechanistic Detail: Allows for competition, supershift, and mutagenesis experiments to define sequence specificity and protein complex composition.
- Speed and Cost-Effectiveness: Rapid, low-cost validation of specific interactions.
Primary Limitations vs. ChIP-seq:
- In Vitro Context: Lacks chromatin and cellular context; binding may not reflect in vivo reality.
- Low-Throughput: Examines one sequence at a time, unlike genome-wide ChIP-seq.
- Requires Purified Components: Needs pure, active protein and defined oligonucleotides.
Detailed Experimental Protocol
Standard Radioactive EMSA Protocol
A. Probe Preparation (Radiolabeling)
- End-Labeling: Incubate 1-10 pmol of dsDNA oligonucleotide with T4 Polynucleotide Kinase (PNK) and [γ-³²P]ATP in 1X PNK buffer for 30 minutes at 37°C.
- Purification: Remove unincorporated nucleotides using a spin column (e.g., Sephadex G-25) or ethanol precipitation.
- Quantification: Measure radioactivity with a scintillation counter. Aim for >50,000 cpm/µL specific activity.
B. Binding Reaction
- Prepare a 10-20 µL reaction mix on ice:
- 1X Binding Buffer (typically: 10 mM Tris, 50 mM KCl, 1 mM DTT, 2.5% Glycerol, 0.05% NP-40, pH 7.5).
- 1 µg Poly(dI-dC) or other non-specific competitor DNA.
- 0.1-1 ng labeled DNA probe (~10,000 cpm).
- Purified protein or nuclear extract (vary amount for titration).
- Incubate at room temperature or 4°C for 20-30 minutes.
C. Electrophoresis and Detection
- Pre-run a 4-10% non-denaturing polyacrylamide gel (29:1 acrylamide:bis) in 0.5X TBE buffer at 100V for 30-60 minutes at 4°C.
- Load binding reactions mixed with non-denaturing loading dye.
- Run gel at constant voltage (100-150V) until the bromophenol blue dye migrates ⅔ of the gel length. Maintain 4°C.
- Transfer gel to Whatman paper, dry under vacuum, and expose to a phosphorimager screen overnight.
- Scan the screen and analyze band intensity.
Key Experimental Variations
- Competition EMSA: Include 10- to 200-fold molar excess of unlabeled competitor DNA (specific or mutant) in the binding reaction to assess binding specificity.
- Supershift EMSA: Add 1-2 µg of antibody specific to the DNA-binding protein after the initial binding reaction. A further reduction in mobility ("supershift") confirms protein identity.
- Fluorescent/Chromogenic EMSA: Use fluorescently (e.g., Cy5, FAM) or biotin-labeled probes. Detection is via gel imaging systems or streptavidin-HRP conjugate with chemiluminescent substrate, respectively. This eliminates radiation hazards.
Quantitative Data & Comparison to ChIP-seq
Table 1: Quantitative Binding Data Obtainable from EMSA Titration Experiments
| Parameter | Description | Typical EMSA Range | Calculation Method |
|---|---|---|---|
| Apparent Kd (Dissociation Constant) | Protein concentration at which 50% of the probe is bound. Measures binding affinity. | 1 pM - 100 nM | Plot fraction bound vs. log[protein]; fit with Hill or logistic equation. |
| Fraction Bound | Proportion of total probe in complex. | 0 to 1.0 | (Intensity of complex band) / (Intensity of complex + free probe bands). |
| Hill Coefficient (n) | Degree of cooperativity in binding. | ~1 (non-cooperative) to >1 (cooperative) | Slope from Hill plot (log[bound/free] vs. log[protein]). |
Table 2: Strategic Comparison: EMSA vs. ChIP-seq for TF Binding Studies
| Aspect | EMSA (In Vitro) | ChIP-seq (In Vivo) |
|---|---|---|
| Primary Objective | Prove direct binding & quantify biophysical parameters. | Map genomic binding locations in a cellular context. |
| Throughput | Low (single sequence/condition). | High (genome-wide). |
| Context | Reduced system; purified components. | Native chromatin & cellular environment. |
| Quantitative Output | Affinity (Kd), stoichiometry, cooperativity. | Relative enrichment peaks; qualitative/relative occupancy. |
| Key Requirement | Purified, active protein; known DNA sequence. | Specific antibody; viable cells. |
| Time to Result | 1-2 days. | 3-7 days. |
| Optimal Use Case | Mechanistic validation of specific ChIP-seq hits, mutational analysis, co-factor requirement. | Discovery of novel binding sites, genomic context, epigenetic state correlation. |
Visualizing EMSA Principles and Workflows
The Scientist's Toolkit: Essential Research Reagents for EMSA
Table 3: Key Research Reagent Solutions for EMSA
| Reagent/Material | Function & Purpose | Key Considerations |
|---|---|---|
| Purified Protein | The DNA-binding protein of interest. Source: recombinant or purified from native tissue. | Requires functional activity. Purity affects specificity; tags (His, GST) should not interfere with DNA binding. |
| Labeled DNA Probe | The target DNA sequence (typically 20-40 bp dsDNA). Label: ³²P, Fluorescent dye (Cy5), or Biotin. | Must contain the suspected protein binding motif. High specific activity/signal is critical for detection. |
| Non-Specific Competitor DNA | e.g., Poly(dI-dC), sheared salmon sperm DNA, or non-specific oligonucleotides. | Suppresses non-sequence-specific protein binding to the probe. Type and amount must be optimized. |
| Binding Buffer | Provides optimal ionic strength, pH, and stabilizing agents (glycerol, DTT, NP-40) for the interaction. | Conditions (Mg²⁺, Zn²⁺, etc.) must be optimized for each protein-DNA pair. |
| Non-Denaturing Gel Matrix | Typically 4-10% polyacrylamide (29:1 or 37.5:1 acrylamide:bis) or agarose. | Separates complex from free probe based on size/charge/shape. Acrylamide offers higher resolution for small probes. |
| Electrophoresis Buffer | 0.5X Tris-Borate-EDTA (TBE) or Tris-Glycine. Low ionic strength maintains complex stability during run. | Often pre-chilled and run at 4°C to stabilize weaker complexes. |
| Detection System | Phosphorimager (³²P), fluorescence scanner, or chemiluminescence imager (Biotin). | Choice dictates probe labeling strategy. Sensitivity and dynamic range vary. |
| Specific & Mutant Competitor Oligos | Unlabeled oligonucleotides identical to the probe (specific) or with mutations in the binding site. | Essential for demonstrating binding sequence specificity in competition assays. |
| Antibody for Supershift | Antibody specific to the DNA-binding protein or an associated tag. | Confirms protein identity in the complex. Must not disrupt the protein-DNA interaction. |
The study of transcription factor (TF)-DNA interactions is fundamental to understanding gene regulation. Two predominant techniques for this are Electrophoretic Mobility Shift Assay (EMSA) and Chromatin Immunoprecipitation followed by sequencing (ChIP-seq). While EMSA provides a powerful in vitro method to confirm direct binding and assess binding affinity using purified components, it lacks the physiological context of living cells. This whitepaper details the core principle of ChIP-seq, which addresses this critical gap by enabling the in vivo mapping of protein-DNA interactions within their native chromatin landscape. The broader thesis argues that ChIP-seq and EMSA are complementary: EMSA offers biochemical precision under controlled conditions, whereas ChIP-seq delivers a genome-wide, functional snapshot of binding events in their natural cellular environment, making it indispensable for drug development targeting dysregulated transcriptional programs.
Core Principle and Workflow
The core principle of ChIP-seq is to cross-link proteins to DNA in vivo, selectively immunoprecipitate the protein-of-interest with its bound DNA fragments, and then identify the associated DNA sequences via high-throughput sequencing. This generates a genome-wide map of binding sites.
Diagram Title: ChIP-seq Experimental Workflow
Detailed Experimental Protocol
Protocol for Native ChIP-seq for Transcription Factors
Day 1: Crosslinking and Cell Lysis
- Crosslinking: Treat cells (~1x10^7) with 1% formaldehyde for 10 minutes at room temperature with gentle agitation to fix protein-DNA interactions.
- Quenching: Add glycine to a final concentration of 0.125 M and incubate for 5 minutes.
- Wash: Wash cells twice with ice-cold PBS. Pellet cells and flash-freeze or proceed.
- Lysis: Resuspend cell pellet in 1 mL Lysis Buffer I (50 mM HEPES-KOH pH 7.5, 140 mM NaCl, 1 mM EDTA, 10% Glycerol, 0.5% NP-40, 0.25% Triton X-100, plus protease inhibitors). Incubate 10 minutes on ice.
- Wash: Pellet nuclei. Resuspend in 1 mL Lysis Buffer II (10 mM Tris-HCl pH 8.0, 200 mM NaCl, 1 mM EDTA, 0.5 mM EGTA, plus protease inhibitors). Incubate 10 minutes on ice. Pellet nuclei.
- Sonication: Resuspend pellet in 1 mL Shearing Buffer (0.1% SDS, 1 mM EDTA, 10 mM Tris-HCl pH 8.0). Sonicate chromatin to an average fragment size of 200-500 bp using a focused ultrasonicator (e.g., Covaris). Optimization is critical.
- Clarification: Centrifuge at 20,000 x g for 10 min at 4°C. Transfer supernatant (sheared chromatin) to a new tube.
Day 2: Immunoprecipitation and Wash
- Pre-clear: Add 50 µL of Protein A/G magnetic beads to chromatin. Rotate for 1 hour at 4°C. Discard beads.
- Immunoprecipitation: Add 1-10 µg of specific antibody (or IgG control) to pre-cleared chromatin. Rotate overnight at 4°C.
- Capture: Add 50 µL of blocked Protein A/G magnetic beads. Rotate for 2 hours at 4°C.
- Wash: Pellet beads and perform sequential 5-minute washes on ice with:
- 1 mL Low Salt Wash Buffer (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris-HCl pH 8.0, 150 mM NaCl).
- 1 mL High Salt Wash Buffer (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris-HCl pH 8.0, 500 mM NaCl).
- 1 mL LiCl Wash Buffer (0.25 M LiCl, 1% NP-40, 1% Na-deoxycholate, 1 mM EDTA, 10 mM Tris-HCl pH 8.0).
- 2 x 1 mL TE Buffer (10 mM Tris-HCl pH 8.0, 1 mM EDTA).
Day 3: Elution and Library Preparation
- Elution: Freshly prepare Elution Buffer (1% SDS, 100 mM NaHCO3). Add 150 µL to beads and incubate at 65°C for 15 minutes with shaking. Collect supernatant. Repeat and combine eluates (~300 µL total).
- Reverse Crosslinks: Add NaCl to a final concentration of 200 mM and RNase A. Incubate at 65°C overnight.
- DNA Purification: Add Proteinase K and incubate at 55°C for 2 hours. Purify DNA using a PCR purification kit (e.g., Qiagen). Elute in 30 µL EB buffer.
- Library Preparation and Sequencing: Use the purified DNA to construct a sequencing library compatible with your platform (e.g., Illumina). This involves end-repair, A-tailing, adapter ligation, and PCR amplification. Validate libraries by Bioanalyzer and quantify by qPCR. Sequence on an appropriate platform (e.g., Illumina NovaSeq).
Data Presentation: ChIP-seq vs. EMSA
Table 1: Quantitative Comparison of ChIP-seq and EMSA Core Characteristics
| Parameter | ChIP-seq | EMSA (Gel Shift) |
|---|---|---|
| Binding Context | In vivo (within native chromatin) | In vitro (purified components) |
| Throughput | Genome-wide (10^4 - 10^5 binding sites per experiment) | Low-throughput (1 - 3 DNA probes per gel) |
| Quantitative Output | Relative enrichment (peak height), binding location | Binding affinity (Kd), stoichiometry |
| Primary Data | Sequence reads mapped to a reference genome | Shifted band intensity on a gel |
| Key Metric | Number of significant peaks (FDR < 0.01); Read density | Dissociation Constant (Kd) in nM/pM range |
| Typical Resolution | 100-300 bp (based on fragment size) | Single binding site resolution (exact sequence probe) |
| Time to Result | 5-7 days (experiment) + extensive bioinformatics analysis | 1-2 days |
| Ability to Detect Indirect Binding | Yes (via other crosslinked proteins) | No (requires direct protein-DNA interaction) |
| Cost per Experiment | High ($1,000 - $3,000+ for sequencing) | Low (< $100 per probe) |
Table 2: Typical ChIP-seq Sequencing and Alignment Metrics
| Metric | Recommended/ Typical Value | Explanation |
|---|---|---|
| Sequencing Depth | 20-40 million reads per sample | Sufficient for transcription factor mapping; more for broad histone marks. |
| Alignment Rate | > 80% | Percentage of reads uniquely mapping to the reference genome. |
| Fraction of Reads in Peaks (FRiP) | 1-5% (TFs), 10-30% (histone marks) | Key quality metric; indicates successful enrichment. |
| Peak Number | Varies widely (1,000 - 50,000) | Depends on TF abundance, antibody quality, and cellular context. |
| Peak Width at Half Maximum | ~200 bp (sharp TF peaks) | Characteristic of point-source binding events. |
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for ChIP-seq Experiments
| Item | Function/Explanation |
|---|---|
| Formaldehyde (37%) | Reversible crosslinker to covalently bind proteins to DNA in vivo. |
| Protease Inhibitor Cocktail | Prevents degradation of the target protein and chromatin-associated factors during lysis and IP. |
| Specific, Validated Antibody | The most critical reagent. Must be ChIP-grade, with proven specificity for the target protein in immunoprecipitation. |
| Protein A/G Magnetic Beads | Efficient capture of antibody-protein-DNA complexes for easy washing and elution. |
| Covaris or Bioruptor | Instrument for consistent, reproducible ultrasonic shearing of chromatin to optimal fragment size. |
| DNA Purification Kit (SPRI) | For efficient cleanup and size selection of DNA after elution and during library preparation. |
| Illumina-Compatible Library Prep Kit | Streamlines conversion of immunoprecipitated DNA into a sequencing-ready library. |
| Control IgG | Isotype-matched non-specific antibody for performing a control IP to assess background noise. |
| Input DNA | A sample of sheared, non-immunoprecipitated chromatin. Serves as control for sequencing and peak calling. |
Pathway: From ChIP-seq Data to Biological Insight
Diagram Title: ChIP-seq Data Analysis Pathway
The study of transcription factor (TF)-DNA interactions is fundamental to understanding gene regulation. Chromatin Immunoprecipitation followed by sequencing (ChIP-seq) and Electrophoretic Mobility Shift Assay (EMSA) represent two principal, complementary methodologies in this field. The efficacy, specificity, and reproducibility of both techniques are critically dependent on their core components: antibodies for target capture, probes for DNA detection, and beads for biomolecular separation. This guide provides a technical deep dive into these reagents, framed within the comparative application of ChIP-seq and EMSA for TF binding research.
Core Components: Technical Specifications and Applications
Antibodies: Specificity is Paramount
Antibodies are the cornerstone of ChIP-seq, used to immunoprecipitate the protein-DNA complex. Their performance dictates the success of the experiment.
- Primary Antibodies: Must be highly specific for the TF of interest and capable of recognizing the protein in its native, cross-linked state. Monoclonal antibodies offer high specificity, whereas high-quality polyclonals may offer higher signal.
- Validation: ChIP-grade validation is essential. This includes demonstration of use in IP, absence of signal in knockout cells, and enrichment at positive control genomic loci.
- EMSA Context: While not always required, antibodies can be used in "supershift" EMSA to confirm TF identity by further retarding the probe-antibody-protein complex.
Probes: The Detectable Target
Probes are nucleic acid sequences used to detect TF binding.
- ChIP-seq: The "probe" is the entire population of sheared, immunoprecipitated genomic DNA, which is converted into a sequencing library. Adapters containing sequencing primer sites and barcodes are ligated.
- EMSA: Probes are short (20-40 bp), double-stranded DNA oligonucleotides containing the suspected TF binding motif. They are typically labeled with fluorophores, biotin, or radioisotopes (³²P) for detection. Cold (unlabeled) probes are used in competition experiments.
Beads: The Workhorse for Isolation
Beads provide a solid-phase matrix for separation.
- Magnetic Beads: The current standard for both techniques due to ease of use and reduced background.
- Protein A/G Beads: Used in ChIP-seq to capture antibody-TF-DNA complexes. Protein A and G have different binding affinities for antibody Fc regions across species and subclasses.
- Streptavidin Beads: Often used in EMSA or biotinylated probe-based ChIP variants to capture biotin-labeled DNA-protein complexes.
- Blocking: Beads must be thoroughly blocked (e.g., with BSA, salmon sperm DNA) to prevent non-specific binding of DNA or proteins.
Quantitative Comparison: ChIP-seq vs EMSA Reagent Requirements
Table 1: Core Reagent Comparison Between ChIP-seq and EMSA
| Component | ChIP-seq | EMSA | Primary Function |
|---|---|---|---|
| Antibody | Mandatory. ChIP-grade, high specificity. | Optional (for supershift). | Isolate specific TF-DNA complexes (ChIP). Identify TF (EMSA supershift). |
| Probe | Entire genomic library (~200-300 bp fragments). | Single, short, defined dsDNA oligo (20-40 bp). | Provide template for sequencing. Serve as labeled target for in vitro binding. |
| Label | Sequencing adapters (Illumina indexes). | Fluorophore, Biotin, or ³²P. | Enable multiplexed NGS. Enable gel visualization. |
| Beads | Magnetic Protein A/G. | Typically none (gel electrophoresis). Streptavidin for pull-down variants. | Solid-phase IP separation. Capture biotinylated complexes. |
| Throughput | Genome-wide, high-throughput. | Low-throughput, single-locus. | |
| Binding Context | In vivo, native chromatin. | In vitro, naked DNA. |
Table 2: Typical Experimental Input Requirements
| Parameter | Standard ChIP-seq | Standard EMSA |
|---|---|---|
| Cells per IP | 0.5 - 5 x 10⁶ | N/A |
| Nuclear Extract | N/A | 2 - 10 µg |
| Antibody per rxn | 1 - 5 µg | 0.5 - 2 µg (supershift) |
| Labeled Probe | N/A | 0.1 - 1 pmol |
| Assay Time | 2-4 days | 4-6 hours |
Detailed Methodologies
Protocol: Native ChIP-seq for a Transcription Factor
Day 1: Cell Crosslinking & Lysis
- Crosslink proteins to DNA in cultured cells using 1% formaldehyde for 10 min at RT. Quench with glycine.
- Harvest cells, wash with cold PBS. Lyse cells in SDS Lysis Buffer (1% SDS, 10 mM EDTA, 50 mM Tris-HCl pH 8.1) with protease inhibitors.
- Sonicate chromatin to shear DNA to 200-500 bp fragments. Verify fragment size by agarose gel electrophoresis.
- Dilute lysate 10-fold in ChIP Dilution Buffer (0.01% SDS, 1.1% Triton X-100, 1.2 mM EDTA, 16.7 mM Tris-HCl pH 8.1, 167 mM NaCl).
Day 1: Immunoprecipitation
- Pre-clear lysate with Protein A/G Magnetic Beads for 1 hour at 4°C.
- Incubate supernatant with 2-5 µg of target-specific antibody (or species-matched IgG control) overnight at 4°C with rotation.
Day 2: Bead Capture & Washes
- Add 50 µL of pre-blocked Protein A/G Magnetic Beads. Incubate 2 hours at 4°C.
- Capture beads on a magnet. Wash sequentially for 5 min each with:
- Low Salt Wash Buffer (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris-HCl pH 8.1, 150 mM NaCl).
- High Salt Wash Buffer (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris-HCl pH 8.1, 500 mM NaCl).
- LiCl Wash Buffer (0.25 M LiCl, 1% NP-40, 1% Na-deoxycholate, 1 mM EDTA, 10 mM Tris-HCl pH 8.1).
- Two washes with TE Buffer (10 mM Tris-HCl pH 8.0, 1 mM EDTA).
Day 2: Elution & Decrosslinking
- Elute complexes twice with 100 µL Fresh Elution Buffer (1% SDS, 0.1 M NaHCO₃). Combine eluates.
- Add NaCl to 200 mM final and reverse crosslinks by heating at 65°C overnight.
Day 3: DNA Purification & Library Prep
- Treat with RNase A and Proteinase K. Purify DNA using a silica-column-based kit.
- Quantify DNA (e.g., by Qubit). Use 1-10 ng of ChIP DNA for standard library preparation (end-repair, A-tailing, adapter ligation, PCR amplification).
Protocol: Non-Radioactive EMSA with Supershift
Part A: Probe Preparation
- Anneal complementary single-stranded oligonucleotides containing the TF binding site: Mix in equimolar ratio (e.g., 100 µM each) in annealing buffer (10 mM Tris, 50 mM NaCl, 1 mM EDTA, pH 7.5-8.0). Heat to 95°C for 5 min, cool slowly to RT.
- Label 100 ng of dsDNA probe at the 3' end using a Biotin 3'-End Labeling Kit. Purify labeled probe.
Part B: Binding Reaction & Electrophoresis
- Prepare 20 µL binding reactions on ice:
- 1X Binding Buffer (10 mM Tris, 50 mM KCl, 1 mM DTT, 2.5% Glycerol, 5 mM MgCl₂, 0.05% NP-40, pH 7.5).
- 1 µg poly(dI·dC) as non-specific competitor.
- 2-5 µg nuclear protein extract.
- (Optional for competition): 100-fold molar excess of unlabeled probe.
- (Optional for supershift): 0.5-2 µg of antibody.
- Incubate 20 min at RT.
- Add 0.5-1 pmol of biotinylated probe. Incubate 20 min at RT.
- Load reaction onto a pre-run 6% non-denaturing polyacrylamide gel in 0.5X TBE buffer. Run at 100 V for 60-90 min at 4°C.
Part C: Transfer & Detection
- Electroblot DNA to a positively charged nylon membrane.
- Crosslink DNA to membrane using UV light.
- Detect biotinylated probe using a Chemiluminescent Nucleic Acid Detection Kit and imaging.
Visualizations
Diagram 1: ChIP-seq vs EMSA Workflow Comparison
Diagram 2: Reagent Role in Complex Formation & Detection
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Reagents for TF Binding Studies
| Reagent Category | Specific Example | Function & Critical Notes |
|---|---|---|
| ChIP-seq Antibodies | Anti-RNA Polymerase II (CTD repeat), Anti-H3K27ac, TF-specific (e.g., Anti-p65). | Positive control (Pol II, H3K27ac) validates protocol. Target antibody must be ChIP-grade. |
| EMSA Probes | Biotin- or Cy5-labeled dsDNA oligo containing consensus AP-1 site. | Provides detectable target for in vitro binding. Consensus sites serve as positive controls. |
| Magnetic Beads | Dynabeads Protein A/G, Streptavidin M-280. | Solid-phase separation. Block thoroughly with BSA/non-specific DNA. |
| Crosslinker | Formaldehyde (37% solution), DSG (for distal crosslinking). | Captures transient in vivo interactions. Quenching is critical. |
| Sonication System | Covaris focused ultrasonicator, Bioruptor. | Shears chromatin to optimal size (200-500 bp). Must be standardized. |
| Non-Specific Competitor | Poly(dI·dC), Salmon Sperm DNA. | Reduces non-specific protein-DNA binding in EMSA and ChIP. |
| Library Prep Kit | Illumina TruSeq ChIP Library Prep Kit, NEB Next Ultra II. | Converts immunoprecipitated DNA into sequencer-compatible libraries. |
| Detection for EMSA | Chemiluminescent Nucleic Acid Detection Kit (e.g., Thermo Scientific LightShift). | Enables sensitive, non-radioactive detection of biotinylated probes. |
This whitepaper addresses the critical distinction between in vitro protein-nucleic acid interaction and in vivo genomic occupancy, a core concept in transcriptional regulation research. Framed within the comparative analysis of Chromatin Immunoprecipitation followed by sequencing (ChIP-seq) and Electrophoretic Mobility Shift Assay (EMSA), we dissect the technical and biological factors that lead to divergent findings between these foundational methods.
Core Conceptual Distinction
In vitro binding, typified by EMSA, measures the biophysical potential for a transcription factor (TF) to interact with a naked DNA sequence. Genomic occupancy, measured by ChIP-seq, identifies where a TF is bound within the native chromatin landscape of a living cell. The discrepancy between these contexts—often termed "binding vs. occupancy"—is driven by chromatin accessibility, co-factors, DNA methylation, and cellular signaling.
Quantitative Comparison of ChIP-seq vs. EMSA
Table 1: Methodological and Output Comparison
| Aspect | EMSA (In Vitro Binding) | ChIP-seq (Genomic Occupancy) |
|---|---|---|
| System | Cell-free, purified components | Intact cells/nuclei, native chromatin |
| Throughput | Low (single probe per assay) | High (genome-wide) |
| Key Readout | Binding affinity/potential (Kd) | Occupancy location & intensity (peak calls) |
| Primary Output | Retardation band on gel | Sequencing reads mapped to genome |
| Identifies | Canonical binding motif | Functional regulatory elements (enhancers, promoters) |
| Influenced by Chromatin | No | Yes (critical confounder) |
| Typical Resolution | Binding site within probe (~10-30 bp) | 100-300 bp (from sheared chromatin) |
| False Positives | Non-specific protein-DNA interactions | Antibody non-specificity, open chromatin artifacts |
| False Negatives | Misses chromatin-dependent binding | Misses low-affinity/transient binding |
Table 2: Concordance Analysis (Representative Data)
| Study Context | % EMSA-validated motifs found in ChIP-seq peaks | % ChIP-seq peaks containing EMSA-validated motif | Key Determinant of Discordance |
|---|---|---|---|
| Pioneer TFs | ~15-30% | ~60-80% | Chromatin remodeling capacity |
| Non-pioneer TFs | ~40-70% | ~20-50% | Pre-existing chromatin accessibility (ATAC-seq signal) |
| Inducible TFs (e.g., NF-κB) | >90% (post-stimulus) | ~70-90% (post-stimulus) | Cellular signaling state & nuclear translocation |
Detailed Experimental Protocols
Protocol 1: Electrophoretic Mobility Shift Assay (EMSA)
Objective: To detect in vitro interaction between a purified transcription factor and a radiolabeled DNA probe containing a putative binding motif.
Key Reagents & Solutions:
- Binding Buffer (10X): 100 mM Tris, 500 mM KCl, 10 mM DTT, pH 7.5. Stabilizes protein-DNA interaction.
- Poly(dI:dC): Non-specific competitor DNA to suppress protein binding to non-specific sequences.
- ³²P-end-labeled DNA probe: 20-30 bp oligonucleotide containing consensus motif.
- Purified TF protein: Full-length or DNA-binding domain (DBD) recombinant protein.
- Non-radiolabeled competitor probe: For specificity validation (cold competition).
- Mutation competitor probe: Probe with mutated motif to confirm binding specificity.
- Anti-TF antibody: For supershift assay confirmation.
Procedure:
- Probe Labeling: Label 10-50 ng of double-stranded DNA probe with [γ-³²P]ATP using T4 Polynucleotide Kinase. Purify using a spin column.
- Binding Reaction: In a 20 µL volume, combine:
- 2 µL 10X Binding Buffer
- 1 µg Poly(dI:dC)
- 1-10 µg nuclear extract or 10-200 ng purified TF protein
- Radiolabeled probe (~20,000 cpm)
- (Optional) 50-200-fold molar excess of cold competitor probe.
- Incubate 20-30 minutes at room temperature.
- Electrophoresis: Load reaction onto a pre-run, non-denaturing 4-6% polyacrylamide gel in 0.5X TBE buffer. Run at 100-150V at 4°C until bromophenol blue dye nears bottom.
- Detection: Dry gel and expose to a phosphorimager screen overnight. Visualize shifted protein-DNA complex bands.
Protocol 2: Chromatin Immunoprecipitation Sequencing (ChIP-seq)
Objective: To map genome-wide occupancy of a transcription factor in its native chromatin context.
Key Reagents & Solutions:
- Crosslinking Reagent: 1% Formaldehyde. Fixes protein-DNA interactions in living cells.
- Sonication Shearing Buffer: 1% SDS, 10 mM EDTA, 50 mM Tris-HCl, pH 8.1. Facilitates chromatin fragmentation.
- ChIP-Grade Anti-TF Antibody: Validated for specificity and efficiency in IP.
- Protein A/G Magnetic Beads: For antibody-immunocomplex capture.
- ChIP Elution Buffer: 1% SDS, 0.1M NaHCO₃.
- DNA Clean-up Columns: For purifying immunoprecipitated DNA.
- Library Preparation Kit: For next-generation sequencing (NGS) adapter ligation and amplification.
Procedure:
- Crosslinking & Quenching: Treat cells with 1% formaldehyde for 10 min at RT. Quench with 125 mM glycine.
- Cell Lysis & Sonication: Lyse cells, isolate nuclei. Sonicate chromatin to 200-500 bp fragments using a focused ultrasonicator. Verify fragment size by agarose gel electrophoresis.
- Immunoprecipitation: Pre-clear chromatin lysate with beads. Incubate lysate with specific antibody overnight at 4°C. Add Protein A/G beads for 2 hours. Wash beads with low-salt, high-salt, LiCl, and TE buffers.
- Elution & Reverse Crosslinking: Elute complexes in elution buffer. Add 5M NaCl and reverse crosslinks at 65°C overnight.
- DNA Purification: Treat with RNase A and Proteinase K. Purify DNA using a spin column.
- Library Prep & Sequencing: Prepare sequencing library from ChIP DNA and an input DNA control. Sequence on an NGS platform (e.g., Illumina).
- Bioinformatics Analysis: Map reads to reference genome, call peaks (e.g., using MACS2), and perform motif enrichment analysis.
Visualizing the Workflow and Biological Context
Title: EMSA Experimental Workflow
Title: ChIP-seq Experimental Workflow
Title: Factors Defining In Vitro vs Genomic Binding
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for TF Binding Studies
| Item | Function & Relevance | Example/Note |
|---|---|---|
| ChIP-Validated Antibodies | High-specificity antibody for immunoprecipitating native TF-chromatin complexes. Critical for ChIP-seq success. | Must be validated for application; check databases like CiteAb. |
| Recombinant TF Protein | Purified, active TF for in vitro assays (EMSA, SELEX, SPR) to define intrinsic binding properties. | Often tagged (GST, His) for purification. Full-length vs DBD. |
| Magnetic Protein A/G Beads | Efficient capture of antibody-TF-chromatin complexes for ChIP, reducing background. | Superior to agarose beads for washing efficiency. |
| Next-Gen Sequencing Library Prep Kit | Prepares immunoprecipitated DNA for sequencing; key for low-input ChIP-seq. | Kits optimized for low DNA input (e.g., ThruPLEX). |
| Validated Consensus & Mutant Oligonucleotides | Probes for EMSA competition controls and motif validation. | Critical for establishing binding specificity in vitro. |
| Chromatin Shearing Reagents & Equipment | Consistent fragmentation of crosslinked chromatin to optimal size (200-500 bp). | Focused ultrasonicator (Covaris) or enzymatic shearing kit. |
| Cell Line with Endogenous Tag (e.g., dTAG) | Enables precise depletion or study of TFs without reliance on antibodies. | Genetic knock-in system for acute protein degradation. |
| ATAC-seq Kit | Maps open chromatin regions in parallel to ChIP-seq to distinguish accessibility-driven occupancy. | Essential for interpreting ChIP-negative/EMSA-positive results. |
EMSA defines the fundamental, biophysical binding grammar of a TF, while ChIP-seq reveals the functional, contextual sentence it forms within the genomic narrative. Discrepancies are not methodological failures but insights into biology: a ChIP-seq peak without strong in vitro affinity may indicate co-factor-dependent stabilization, while a strong EMSA site absent in vivo highlights chromatin-mediated repression. The integrated use of both methods, complemented by chromatin accessibility assays (ATAC-seq) and perturbation studies, provides a complete picture of transcriptional regulation, directly informing drug discovery targeting pathological gene programs.
Primary Applications in Transcription Factor Research
Within the framework of a thesis comparing Chromatin Immunoprecipitation Sequencing (ChIP-seq) and Electrophoretic Mobility Shift Assay (EMSA) for transcription factor (TF) binding research, understanding their primary applications is crucial. These techniques address distinct but complementary questions in gene regulation. This guide details their core technical applications, protocols, and data interpretation, providing researchers and drug development professionals with a foundation for experimental design.
Table 1: Primary Applications of ChIP-seq vs. EMSA in TF Research
| Application Dimension | ChIP-seq | EMSA |
|---|---|---|
| Primary Objective | Genome-wide mapping of in vivo TF binding sites. | Detection of in vitro protein-nucleic acid interactions. |
| Binding Context | Native chromatin environment within cells. | Purified components in a cell-free system. |
| Throughput & Scale | High-throughput, maps thousands of sites genome-wide. | Low-throughput, analyzes single or few DNA sequences per assay. |
| Quantitative Output | Semi-quantitative (enrichment peaks). Can measure differential binding. | Semi-quantitative (band shift intensity). Can calculate binding affinity (Kd). |
| Key Resolved Information | Genomic location, sequence motif, co-binding patterns, correlation with gene expression. | Confirmation of direct binding, sequence specificity, complex stoichiometry. |
| Typical Use Case | Discovering novel TF targets, defining regulatory networks, integration with epigenomics. | Validating direct TF-DNA interaction, mapping minimal binding motif, testing mutant probes. |
Detailed Methodologies
Protocol 1: Standard ChIP-seq for Transcription Factors
Objective: To identify genome-wide binding sites of a transcription factor in its native cellular context. Key Steps:
- Crosslinking: Treat cells (~1x10^7) with 1% formaldehyde for 10 min at room temperature to fix protein-DNA interactions. Quench with 125mM glycine.
- Cell Lysis & Chromatin Preparation: Lyse cells, isolate nuclei, and shear chromatin via sonication to fragment sizes of 200-500 bp.
- Immunoprecipitation: Incubate sheared chromatin with a high-specificity antibody against the target TF (e.g., 1-10 µg per reaction) overnight at 4°C. Use Protein A/G magnetic beads for capture.
- Washing & Elution: Wash beads stringently (e.g., low salt, high salt, LiCl, TE buffers). Elute bound complexes with freshly prepared elution buffer (1% SDS, 0.1M NaHCO3).
- Reverse Crosslinking & Purification: Incubate eluates at 65°C overnight with NaCl to reverse crosslinks. Treat with Proteinase K and RNase A. Purify DNA using spin columns.
- Library Prep & Sequencing: Prepare sequencing library from ChIP-DNA (and Input DNA control) using adaptor ligation and PCR amplification. Sequence on an Illumina platform (typically 20-50 million reads).
Diagram 1: ChIP-seq experimental workflow for TF binding mapping.
Protocol 2: EMSA for TF-DNA Interaction Validation
Objective: To confirm direct, sequence-specific binding of a purified or in vitro translated TF to a target DNA sequence. Key Steps:
- Probe Preparation: Anneal complementary oligonucleotides containing the putative TF binding site. Label the probe at the 5' end with [γ-³²P] ATP using T4 Polynucleotide Kinase. Purify using a spin column.
- Protein Preparation: Use purified recombinant TF protein or nuclear extract containing the TF.
- Binding Reaction: Incubate 10-20 fmol of labeled probe with TF protein (varying amounts) in a binding buffer (10mM HEPES, 50mM KCl, 1mM DTT, 2.5% glycerol, 0.05% NP-40, 50 ng/µL poly(dI•dC)) for 20-30 min at room temperature.
- Electrophoresis: Load the reaction mixture onto a pre-run, non-denaturing polyacrylamide gel (4-6%) in 0.5x TBE buffer. Run at 100-150 V at 4°C to maintain complex stability.
- Detection & Analysis: Transfer gel to filter paper, dry, and expose to a phosphorimager screen or X-ray film. The shifted band (protein-DNA complex) migrates slower than the free probe.
- Competition Assay (Specificity Control): Include a 50-100x molar excess of unlabeled wild-type (competitor) or mutated (non-competitor) oligonucleotide in the binding reaction prior to adding the labeled probe.
Diagram 2: EMSA workflow for validating direct TF-DNA interaction.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for TF Binding Studies
| Item | Function & Application |
|---|---|
| High-Quality TF-specific Antibody | Critical for ChIP-seq specificity. Must be validated for immunoprecipitation (ChIP-grade). |
| Formaldehyde (37%) | Reversible crosslinker for in vivo fixation of TF-DNA interactions in ChIP. |
| Magnetic Protein A/G Beads | Solid-phase support for antibody capture in ChIP, enabling efficient washing. |
| Sonication Device (e.g., Bioruptor) | For consistent chromatin shearing to optimal fragment size in ChIP-seq. |
| Poly(dI•dC) | Non-specific competitor DNA used in EMSA to reduce protein binding to non-target sequences. |
| [γ-³²P] ATP or Chemiluminescent Labeling Kit | For sensitive radioactive or non-radioactive end-labeling of DNA probes in EMSA. |
| Recombinant TF Protein | Purified protein for EMSA, allows study of direct binding without confounding cellular factors. |
| Non-Denaturing PAGE Gel System | For separation of protein-DNA complexes from free probe based on size & charge in EMSA. |
| ChIP-seq Library Prep Kit | Optimized reagents for efficient conversion of low-input ChIP DNA into sequencing libraries. |
| Validated Consensus Oligonucleotides | Positive control probes (e.g., SP1, NF-κB sites) for EMSA optimization. |
Data Interpretation & Integration
Table 3: Quantitative Metrics and Their Interpretation
| Technique | Key Metric | Typical Value/Range | Biological Interpretation |
|---|---|---|---|
| ChIP-seq | Number of Significant Peaks | TF-dependent; from 100s to 100,000s. | Indicates the scope of the TF's genomic occupancy. |
| ChIP-seq | Peak Enrichment (Fold-change over input) | Often 5-fold to >100-fold at high-affinity sites. | Reflects relative binding strength or antibody efficiency. |
| ChIP-seq | Distance from Peak Summit to TSS | Many TFs peak within ±1 kb of TSS. | Suggests direct transcriptional regulatory function. |
| EMSA | Apparent Dissociation Constant (Kd) | nM range (e.g., 1-50 nM). | Quantifies in vitro binding affinity of TF for the probe. |
| EMSA | % of Probe Shifted | Varies with protein concentration; up to >80%. | Estimates fraction of DNA bound under given conditions. |
| EMSA | Competition IC₅₀ | Molar excess needed (e.g., 10-100x). | Measures specificity and relative affinity of competitor DNA. |
ChIP-seq and EMSA serve as foundational pillars in transcription factor research, addressing in vivo binding landscapes and in vitro biochemical mechanisms, respectively. A robust thesis leverages their complementary nature: ChIP-seq generates genome-wide hypotheses on TF occupancy, while EMSA provides mechanistic validation of direct, sequence-specific binding. The choice of technique is dictated by the biological question, ranging from systems-level network discovery to reductionist molecular validation, both essential for advancing therapeutic targeting of TFs.
From Bench to Data: Step-by-Step Protocols and Strategic Applications
In the study of transcription factor (TF)-DNA interactions, researchers often choose between in vivo methods like Chromatin Immunoprecipitation followed by sequencing (ChIP-seq) and in vitro techniques such as the Electrophoretic Mobility Shift Assay (EMSA). ChIP-seq provides a genome-wide map of TF binding sites within a native chromatin context, revealing functional regulatory elements in living cells. In contrast, EMSA offers a direct, biochemical validation of specific protein-DNA interactions, allowing for the quantification of binding affinity, kinetics, and complex composition under controlled conditions. This whitepaper provides an in-depth technical guide to EMSA, a critical orthogonal technique for validating ChIP-seq findings and performing detailed mechanistic studies.
Core Principles of EMSA
EMSA exploits the principle that a protein bound to a nucleic acid probe (typically DNA) retards its electrophoretic mobility through a non-denaturing polyacrylamide gel. The shift in migration is visualized, confirming a binding event.
Probe Design and Preparation
The DNA probe is a critical component. It typically contains the suspected TF binding site (cis-element).
Design Guidelines:
- Length: 20-50 base pairs. Longer probes may have secondary structures; shorter ones may not provide sufficient flanking sequence for stable binding.
- Sequence: Center the putative binding motif. Include 10-15 bp of flanking sequence on each side, derived from the native genomic context if possible.
- Labeling: Probes are end-labeled with γ-³²P ATP (radioactive) or biotin/fluorophores (non-radioactive) for detection.
- Control Probes: Always design:
- Mutant Probe: Contains a scrambled or mutated core binding motif.
- Unlabeled Competitor Probe: Identical to the labeled probe, used in competition experiments.
Protocol: Annealing and Labeling of Oligonucleotides
- Annealing: Combine complementary single-stranded oligonucleotides in equimolar ratios in annealing buffer (10 mM Tris, 50 mM NaCl, 1 mM EDTA, pH 8.0). Heat to 95°C for 5 min, then slowly cool to room temperature.
- End-Labeling (Radioactive, T4 PNK):
- Combine: 1 µL double-stranded probe (10 pmol/µL), 2 µL 10x T4 Polynucleotide Kinase (PNK) buffer, 5 µL γ-³²P ATP (3000 Ci/mmol), 1 µL T4 PNK (10 U/µL), 11 µL nuclease-free water.
- Incubate at 37°C for 30 min.
- Purify using a spin column or gel filtration to remove unincorporated nucleotides.
Binding Reaction
The reaction establishes optimal conditions for the specific TF-DNA interaction.
Key Optimization Parameters: pH, ionic strength (KCl/NaCl concentration), presence of divalent cations (Mg²⁺), non-specific competitors (poly(dI:dC)), carrier proteins (BSA), and non-ionic detergents.
Standard Binding Reaction Protocol:
- Prepare a master mix on ice for n+1 reactions. A typical 20 µL reaction contains:
- 2 µL 10x Binding Buffer (100 mM Tris, 500 mM KCl, 10 mM DTT, pH 7.5)
- 1 µL Poly(dI:dC) (1 µg/µL, to mask non-specific binding)
- 1 µL BSA (10 µg/µL, stabilizes protein)
- 1 µL NP-40 (10%, reduces adsorption)
- X µL Nuclear extract or purified protein (2-10 µg)
- Nuclease-free water to 18 µL
- Pre-incubate the master mix (without probe) on ice for 10 min.
- Add 2 µL of labeled probe (approx. 20 fmol).
- Incubate at room temperature (or specific temperature) for 20-30 min.
- For competition assays, add a 50-200 fold molar excess of unlabeled competitor probe (wild-type or mutant) to the master mix before adding the labeled probe.
Table 1: Common EMSA Reaction Components and Their Functions
| Component | Typical Concentration | Function |
|---|---|---|
| Binding Buffer | 1x | Maintains pH and ionic strength optimal for specific binding. |
| Poly(dI:dC) | 0.5-2 µg/µL | Inert polymer that competes for non-specific protein-DNA interactions. |
| BSA | 0.5-1 µg/µL | Carrier protein that stabilizes the TF and prevents adhesion to tubes. |
| DTT | 0.5-1 mM | Reducing agent that maintains protein sulfhydryl groups. |
| MgCl₂ | 0-5 mM | May be required for the DNA-binding fold of some TFs (e.g., zinc fingers). |
| NP-40 / Tween-20 | 0.1% | Non-ionic detergent reduces non-specific binding. |
| Labeled Probe | ~1 nM | The target DNA sequence for detection. |
| Nuclear Extract | 2-10 µg | Source of transcription factor protein. |
Gel Electrophoresis and Analysis
The protein-DNA complex is separated from free probe via non-denaturing polyacrylamide gel electrophoresis (PAGE).
Protocol: Non-Denaturing Gel Electrophoresis
- Gel Preparation: Cast a 4-10% polyacrylamide gel (29:1 acrylamide:bis) in 0.5x TBE buffer. A lower percentage gel better resolves large complexes.
- Pre-run: Pre-electrophorese the gel in 0.5x TBE buffer at 100 V for 30-60 min at 4°C to reach temperature equilibrium and remove ammonium persulfate.
- Loading: Add 5 µL of loading dye (glycerol-based, without SDS or bromophenol blue) to each binding reaction. Load the entire sample.
- Run: Run the gel at 100 V (constant) in 0.5x TBE at 4°C until the dye front is near the bottom (~1.5-2 hours).
- Detection:
- Radioactive: Transfer gel to blotting paper, dry, and expose to a phosphorimager screen.
- Biotin: Electroblot to a positively charged nylon membrane, crosslink, and detect with Streptavidin-HRP and chemiluminescence.
Table 2: Troubleshooting Common EMSA Results
| Problem | Potential Cause | Solution |
|---|---|---|
| No shifted band | Insufficient protein; Probe degradation; Incorrect binding conditions. | Titrate protein amount; Check probe integrity; Optimize buffer (K⁺, Mg²⁺). |
| High background/smearing | Non-specific binding; Too much probe; Gel running too warm. | Increase poly(dI:dC); Reduce probe amount; Run gel at 4°C. |
| Multiple shifted bands | Protein degradation; Other proteins binding; Oligomerization. | Use fresh protease inhibitors; Include specific antibody for supershift; Characterize complexes. |
| Poor gel resolution | Wrong gel %; Incorrect buffer; Air bubbles in gel. | Use lower % acrylamide; Use fresh 0.5x TBE; Degas acrylamide solution. |
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in EMSA |
|---|---|
| T4 Polynucleotide Kinase (PNK) | Catalyzes the transfer of a radioactive phosphate from γ-³²P ATP to the 5'-end of DNA for probe labeling. |
| Poly(dI:dC) | A synthetic alternating copolymer used as a non-specific competitor to absorb non-sequence-specific DNA-binding proteins. |
| Protease Inhibitor Cocktail | Added to protein extracts to prevent degradation of the transcription factor of interest. |
| Non-Radiative Labeling Kits (e.g., Biotin, Digoxigenin) | Provide reagents for end-labeling and subsequent chemiluminescent detection, offering a safer alternative to radioactivity. |
| High-Binding Streptavidin-HRP Conjugate | Used with biotinylated probes for highly sensitive chemiluminescent detection on blotted membranes. |
| Super-shift Antibody | An antibody specific to the TF or an epitope tag. Binding to the protein-DNA complex creates an even larger "supershifted" band, confirming TF identity. |
| Non-Denaturing Acrylamide/Bis Solution (29:1) | The matrix for the gel, optimized for separating native protein-nucleic acid complexes based on size and charge. |
| Cold Competitor Oligonucleotides | Unlabeled wild-type and mutant DNA sequences used to demonstrate binding specificity through competition. |
Experimental Workflow and Data Interpretation Logic
Diagram Title: EMSA Experimental and Validation Logic Flow
Complementary Role to ChIP-seq
The following diagram illustrates the complementary relationship between EMSA and ChIP-seq within a TF research pipeline.
Diagram Title: Complementary Roles of ChIP-seq and EMSA in TF Research
Chromatin immunoprecipitation followed by sequencing (ChIP-seq) is the dominant method for genome-wide profiling of transcription factor (TF) binding sites and histone modifications. Within the broader thesis comparing ChIP-seq to the electrophoretic mobility shift assay (EMSA) for TF binding research, ChIP-seq offers unparalleled in vivo resolution and genomic coverage, though with greater technical complexity. This guide details the core protocol.
Crosslinking
Crosslinking covalently binds proteins, including TFs, to their associated DNA. Formaldehyde (typically 1% final concentration) is most common, with a 10-minute incubation at room temperature. For some repressive or large protein complexes, a dual crosslinking approach with agents like DSG (disuccinimidyl glutarate) may be used. The reaction is quenched with glycine.
Table 1: Common Crosslinking Conditions
| Condition | Agent | Concentration | Incubation Time | Primary Use |
|---|---|---|---|---|
| Standard | Formaldehyde | 1% | 8-12 min | Most TFs, histones |
| Dual | DSG + Formaldehyde | 2 mM + 1% | 20-45 min + 10 min | Repressive complexes, challenging TFs |
| Light | Formaldehyde | 0.5-0.75% | 5 min | To preserve fragile epitopes |
Protocol: Harvest cells, resuspend in media/PBS. Add 37% formaldehyde directly to achieve 1%. Incubate 10 min with gentle shaking. Add 2.5M glycine to 0.125M final to quench. Incubate 5 min. Pellet cells, wash 2x with cold PBS. Pellet can be frozen at -80°C.
Sonication (Chromatin Shearing)
Crosslinked chromatin is fragmented via sonication to an optimal size of 200-500 bp. This can be performed using a bath or probe sonicator. Key parameters include duration, power, and pulse settings, which must be empirically determined per cell type and sonicator.
Table 2: Typical Sonication Parameters for Different Platforms
| Platform | Settings | Peak Power | Time | Cycles | Target Fragment Size |
|---|---|---|---|---|---|
| Probe Sonicator | 20% amplitude | ~50 W | 10 x 30s pulses | 10 | 200-500 bp |
| Bath Sonicator (Covaris) | 140W Peak, 5% Duty Factor | 140 W | 8-12 min | N/A | 200-500 bp |
| Bioruptor (Diagenode) | High Power | N/A | 30s ON / 30s OFF | 15-20 | 200-500 bp |
Protocol: Resuspend fixed cell pellet in lysis/sonication buffer (e.g., 1% SDS, 10mM EDTA, 50mM Tris-HCl pH8.1) with protease inhibitors. Sonicate on ice. Centrifuge to remove debris. Analyze fragment size by running 2% of sheared chromatin on a 1.5% agarose gel or Bioanalyzer.
Immunoprecipitation (IP)
Sheared chromatin is incubated with an antibody specific to the target protein. Antibody-chromatin complexes are then isolated using protein A/G beads.
Protocol: Dilute sheared chromatin 10-fold in IP dilution buffer (e.g., 0.01% SDS, 1.1% Triton X-100, 1.2mM EDTA, 16.7mM Tris-HCl pH8.1, 167mM NaCl). Pre-clear with beads for 1-2h. Incubate supernatant with antibody (1-10 µg) overnight at 4°C. Add pre-blocked Protein A/G beads, incubate 2h. Wash beads sequentially with: Low Salt Wash Buffer, High Salt Wash Buffer, LiCl Wash Buffer, and TE Buffer. Elute complexes with fresh elution buffer (1% SDS, 0.1M NaHCO3). Reverse crosslinks at 65°C overnight with high salt (200mM NaCl). Treat with RNase A and Proteinase K. Purify DNA with spin columns or bead-based cleanup.
Library Preparation for Sequencing
The immunoprecipitated DNA is prepared into a sequencing library compatible with platforms like Illumina. This involves end-repair, A-tailing, adapter ligation, and PCR amplification.
Protocol: Starting with 1-10 ng of ChIP DNA. 1) End Repair: Convert overhangs to phosphorylated blunt ends. 2) A-tailing: Add a single 'A' nucleotide to 3' ends. 3) Adapter Ligation: Ligate indexed adapters with a complementary 'T' overhang. 4) Size Selection: Use SPRI beads to select fragments ~200-500 bp. 5) PCR Amplification: Enrich adapter-ligated fragments with 10-18 cycles of PCR. 6) Cleanup & QC: Purify library and assess concentration/fragment size via qPCR and Bioanalyzer/TapeStation.
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function & Explanation |
|---|---|
| Formaldehyde (37%) | Reversible crosslinking agent. Creates protein-DNA and protein-protein bridges to capture transient interactions. |
| Protein A/G Magnetic Beads | Solid support for antibody capture. Magnetic beads simplify washing and elution steps vs. agarose beads. |
| ChIP-Validated Antibody | Critical for specificity. Must be validated for IP application; poor antibody quality is a major failure point. |
| Protease Inhibitor Cocktail | Prevents degradation of proteins/TFs during cell lysis and sonication. |
| SPRI (Solid Phase Reversible Immobilization) Beads | Magnetic beads for size selection and cleanup of DNA during library prep. Based on PEG/NaCl precipitation. |
| TruSeq or NEBNext Ultra II Library Prep Kit | Commercial kits that provide optimized, pre-mixed reagents for all library prep steps. |
| Covaris or Diagenode Bioruptor Sonicator | Provides consistent, controlled acoustic shearing with minimal heat transfer to preserve epitopes. |
| High Sensitivity DNA Bioanalyzer Chip | Microfluidics-based system for precise quantification and size distribution analysis of sheared chromatin and final libraries. |
Visualizing the ChIP-seq Workflow
Diagram Title: Core ChIP-seq Experimental Workflow
Context: ChIP-seq vs. EMSA in TF Research
ChIP-seq's in vivo mapping capability, where binding is captured in its native chromatin context, contrasts sharply with EMSA's in vitro approach using purified proteins and probe DNA. While EMSA is excellent for probing direct binding affinity and kinetics, ChIP-seq reveals the genome-wide binding landscape within a biological system. The protocol above enables this comprehensive view, though its success hinges on the critical steps of crosslinking optimization, efficient sonication, and antibody specificity.
Within the broader methodological debate of ChIP-seq versus EMSA for transcription factor (TF) research, the Electrophoretic Mobility Shift Assay (EMSA) remains a cornerstone technique. While ChIP-seq excels at genome-wide, in vivo binding site discovery, EMSA provides unparalleled in vitro validation of direct, specific protein-nucleic acid interactions and is indispensable for detailed mechanistic studies. This guide details the precise experimental contexts where EMSA is the optimal choice.
Core Applications: Validating Specificity and Mechanism
EMSA is uniquely positioned to answer two fundamental questions that ChIP-seq cannot:
- Direct Interaction Validation: Does the purified protein of interest bind directly to the target DNA/RNA sequence?
- Interaction Dissection: Which specific nucleotides or protein domains are critical for binding?
The following table contrasts the core capabilities of EMSA and ChIP-seq:
Table 1: Strategic Comparison: EMSA vs. ChIP-seq
| Feature | EMSA (Electrophoretic Mobility Shift Assay) | ChIP-seq (Chromatin Immunoprecipitation Sequencing) |
|---|---|---|
| Primary Purpose | Validate direct, specific protein-nucleic acid interactions in vitro. | Map genome-wide protein binding sites in vivo. |
| Throughput | Low to medium (individual probes). | Very high (entire genome). |
| Direct Binding Proof | Yes. Uses purified components. | Indirect. Requires crosslinking and immunoprecipitation. |
| Resolution | Single binding site (~20-30 bp probe). | ~100-200 bp regions from fragmented chromatin. |
| Quantitative Data | Binding affinity (Kd), stoichiometry. | Relative enrichment scores. |
| Best for Mutagenesis | Excellent. Precise assessment of mutant probe binding. | Limited; requires creating mutant cell lines or organisms. |
| Context | Cell-free, controlled conditions. | Native chromatin, cellular context. |
Detailed Experimental Protocols
Protocol 1: Standard EMSA for Specificity Validation
Objective: To confirm a purified TF binds directly to a suspected DNA consensus sequence.
Key Research Reagent Solutions:
- Purified Protein: Recombinant TF (>90% purity). Essential for proving direct binding.
- 32P- or Fluorescently-Labeled Probe: Double-stranded oligonucleotide containing the putative binding site (20-30 bp). Critical for visualization.
- Poly(dI:dC): A nonspecific competitor DNA that reduces background by binding non-specific proteins.
- Specific & Mutant Unlabeled Competitors: Unlabeled oligonucleotides (identical or with scrambled/mutated site) to demonstrate binding specificity.
- Non-denaturing Polyacrylamide Gel (4-6%): Matrix that separates protein-bound (shifted) from free nucleic acid.
- Binding Buffer (10X): Typically contains Tris-HCl (pH 7.5), KCl, MgCl2, DTT, EDTA, and glycerol.
Methodology:
- Probe Preparation: End-label 1 pmol of dsDNA oligonucleotide with [γ-32P]ATP using T4 Polynucleotide Kinase. Purify using a spin column.
- Binding Reaction: In a 20 µL volume, combine:
- 1X Binding Buffer
- 1 µg Poly(dI:dC)
- ~20 fmol labeled probe (~20,000 cpm)
- Purified TF protein (0-100 nM range)
- Nuclease-free water.
- For competition: Add 50-200-fold molar excess of unlabeled competitor DNA.
- Incubation: Mix and incubate at room temperature for 20-30 minutes.
- Electrophoresis: Load samples onto a pre-run 4-6% native polyacrylamide gel in 0.5X TBE buffer. Run at 100V at 4°C for 60-90 minutes.
- Detection: Dry gel and expose to a phosphorimager screen or X-ray film.
Protocol 2: EMSA-based Mutagenesis Study
Objective: To define the critical nucleotides within a TF binding site.
Methodology:
- Design Probes: Synthesize a series of labeled dsDNA probes where 2-3 base pairs within the core consensus are systematically mutated (e.g., transversions, deletions).
- Perform Parallel EMSAs: Run standard binding reactions (as in Protocol 1) using the wild-type and each mutant probe with a fixed, subsaturating concentration of the TF.
- Quantitative Analysis: Use phosphorimager quantification to determine the fraction of probe bound for each sequence. Calculate relative binding affinity compared to wild-type (set at 100%).
- Data Presentation: Results are best summarized in a table and a bar graph.
Table 2: Example EMSA Mutagenesis Data for a Hypothetical TF "X"
| Probe Name | Sequence (Core Site in Bold) | Relative Binding Affinity (%) | Interpretation |
|---|---|---|---|
| WT | 5'-TACGCGTA-3' | 100 ± 5 | Reference sequence. |
| Mut1 | 5'-TAgCGCGA-3' | 12 ± 3 | G2 is critical for binding. |
| Mut2 | 5'-TAaGCGTA-3' | 95 ± 6 | C3 is not essential. |
| Mut3 | 5'-TACcCcTA-3' | 5 ± 2 | G5 and G6 are both critical. |
Visualizing EMSA's Role in the TF Research Workflow
Diagram Title: EMSA's Niche in TF Binding Research
The Scientist's Toolkit: Essential EMSA Reagents
Table 3: Essential Research Reagent Solutions for EMSA
| Item | Function & Rationale |
|---|---|
| High-Purity Recombinant Protein | Mandatory to prove direct binding. Can be tagged (e.g., GST, His) for purification and supershift experiments. |
| Radioactive (³²P) or Chemifluorescent Probes | Provides high sensitivity for detecting low-abundance or low-affinity complexes. Fluorescent dyes (Cy5, FAM) offer safer alternatives. |
| Non-specific Competitor DNA (Poly(dI:dC)) | Blocks non-specific protein-DNA interactions, reducing background and clarifying specific shifted bands. |
| Specific Unlabeled Competitor Oligo | Demonstrates sequence specificity; excess should abolish the shifted band. |
| Antibody for "Supershift" | Antibody against the TF binds to the protein-DNA complex, causing a further mobility shift, confirming protein identity. |
| Non-denaturing PAGE Gel System | The matrix that resolves complexes based on size/charge without disrupting non-covalent interactions. |
| High-Stringency Binding & Gel Buffers | Optimized ionic strength and pH are crucial for maintaining specific interactions during electrophoresis. |
In the integrative analysis of transcription factor binding, EMSA is not a competitor to ChIP-seq but a vital complementary tool. ChIP-seq identifies where in the genome binding occurs in vivo, while EMSA rigorously proves that the binding is direct and defines how at a nucleotide level through mutagenesis. For validating specific interactions, determining binding affinity, and dissecting precise sequence requirements, EMSA remains the definitive in vitro assay of choice.
The selection of an appropriate methodology is foundational to transcription factor (TF) binding research. While the Electrophoretic Mobility Shift Assay (EMSA) is a classical, in vitro technique for validating specific protein-nucleic acid interactions, Chromatin Immunoprecipitation followed by sequencing (ChIP-seq) has emerged as the premier in vivo method for genome-wide discovery and epigenetic profiling. This guide details when and why to choose ChIP-seq, framing it as the essential tool for unbiased, in vivo mapping of protein-DNA interactions and histone modifications across the entire genome, a capability fundamentally absent in EMSA's targeted, candidate-based approach.
Core Applications: When ChIP-seq is Indispensable
ChIP-seq is the method of choice in the following scenarios, which contrast sharply with EMSA's limited scope:
- De Novo Discovery of Binding Sites: Unbiased identification of all genomic loci bound by a TF or enriched for a specific histone mark without prior knowledge of target sequences.
- Epigenetic Landscape Profiling: Genome-wide mapping of histone modifications (e.g., H3K4me3 for promoters, H3K27ac for enhancers), histone variants, or chromatin-regulatory enzymes.
- Comparative & Condition-Specific Profiling: Investigating dynamic changes in the epigenome or TF occupancy across different cellular states, disease conditions, or in response to stimuli.
- Enhancer and Regulatory Element Identification: Defining active, poised, or repressed regulatory regions across the genome.
Comparative Quantitative Data: ChIP-seq vs. EMSA
Table 1: Fundamental Comparison of ChIP-seq and EMSA
| Feature | ChIP-seq | EMSA (Gel Shift) |
|---|---|---|
| Scope | Genome-wide, discovery-driven | Targeted, candidate-driven |
| Throughput | High (millions of loci per experiment) | Low (typically 1 probe per assay) |
| Context | In vivo, within native chromatin | In vitro, using purified components |
| Primary Output | Map of binding/enrichment peaks across the genome | Confirmation of binding to a specific DNA sequence |
| Quantitative Nature | Semi-quantitative (peak height/counts) | Semi-quantitative (band intensity shift) |
| Ability to Detect Co-binding | Indirect, via peak co-localization | Limited, via supershift with another antibody |
| Typical Time to Result | 4-7 days (library prep to data) | 1-2 days |
| Approximate Cost per Sample | $500 - $1500 (sequencing dependent) | $50 - $200 |
Table 2: Typical ChIP-seq Output Metrics from a Modern Experiment
| Metric | Typical Value/Range | Explanation |
|---|---|---|
| Sequencing Depth | 20 - 50 million reads (histones) | Deeper sequencing (e.g., 50-100M reads) is often required for TFs with fewer, sharper peaks. |
| Peak Number (TF) | 10,000 - 80,000 | Varies greatly by TF, cell type, and statistical threshold. |
| Peak Number (Histone Mark) | 50,000 - 200,000 | Broader marks (e.g., H3K9me3) yield fewer, wider peaks than sharp marks (e.g., H3K4me3). |
| Peak Width | 200 - 500 bp (TF), 1,000 - 5,000 bp (histones) | TF peaks are narrow; histone marks are broader. |
| FRiP Score | >1% (TF), >5% (histones) | Fraction of Reads in Peaks; a key quality control metric. |
Detailed ChIP-seq Experimental Protocol
Principle: Crosslink protein to DNA in vivo, shear chromatin, immunoprecipitate with a specific antibody, then sequence the associated DNA fragments.
Key Protocol Steps:
- Crosslinking: Treat cells with 1% formaldehyde for 8-12 minutes at room temperature to covalently link proteins to DNA. Quench with glycine.
- Cell Lysis & Chromatin Preparation: Lyse cells in SDS buffer. Pellet nuclei. Resuspend in sonication buffer.
- Chromatin Shearing: Using a focused ultrasonicator (e.g., Covaris), shear crosslinked chromatin to an average fragment size of 200-500 bp. Critical: Optimize for each cell type.
- Immunoprecipitation: Pre-clear sheared chromatin with protein A/G beads. Incubate overnight at 4°C with a validated, high-specificity antibody against the target protein or histone mark. Capture immune complexes with beads.
- Washes & Elution: Wash beads stringently with a series of buffers (e.g., low salt, high salt, LiCl, TE) to remove non-specific binding. Elute bound complexes with fresh elution buffer (1% SDS, 0.1M NaHCO3).
- Reverse Crosslinking & Purification: Incubate eluate and input control at 65°C overnight with high salt to reverse crosslinks. Treat with RNase A and Proteinase K. Purify DNA using a spin column or bead-based method.
- Library Preparation & Sequencing: Using ~10 ng of purified ChIP-DNA, perform end-repair, A-tailing, adapter ligation, and limited-cycle PCR amplification. Size-select fragments (typically 200-400 bp). Sequence on an Illumina platform (e.g., NovaSeq) to generate at least 20 million single-end 50-bp reads.
Visualizing the ChIP-seq Workflow and Data Analysis Logic
Title: ChIP-seq Experimental and Computational Workflow
Title: Key Pathways in ChIP-seq Data Analysis
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Reagents and Materials for a Successful ChIP-seq Experiment
| Item | Function & Critical Consideration |
|---|---|
| Validated ChIP-grade Antibody | The most critical reagent. Must be validated for immunoprecipitation under cross-linked conditions. Use ChIP-seq citation databases. |
| Magnetic Protein A/G Beads | For efficient capture of antibody-antigen complexes. Offer low background and ease of handling over agarose beads. |
| Formaldehyde (37%) | Standard crosslinking agent for fixing protein-DNA interactions. Must be fresh for efficient crosslinking. |
| Protease/Phosphatase Inhibitor Cocktails | Essential to prevent degradation and modification of epitopes and chromatin during processing. |
| Covaris or Bioruptor Sonicator | Provides consistent, controllable chromatin shearing to achieve ideal fragment size distribution. |
| DNA Clean & Concentrator Kit | For efficient purification of low-yield ChIP-DNA after reverse crosslinking. |
| High-Sensitivity DNA Assay Kit | Accurate quantification of minute amounts of ChIP-DNA (e.g., Qubit dsDNA HS Assay) is mandatory for library prep. |
| Illumina-Compatible Library Prep Kit | Optimized for low-input DNA, includes all enzymes and buffers for end-prep, ligation, and indexing PCR. |
| Size Selection Beads | SPRI/AMPure XP beads are used to select library fragments in the desired size range, removing adaptor dimers and large fragments. |
| Control Antibodies | Anti-IgG (negative control) and anti-RNA Pol II or a known histone mark (e.g., H3K4me3) as a positive control. |
| Input DNA | A sample of sheared, reverse-crosslinked chromatin saved prior to IP. Serves as the essential control for peak calling. |
In the context of a broader thesis on transcription factor (TF) binding research, a persistent debate centers on the use of in vivo chromatin immunoprecipitation sequencing (ChIP-seq) versus in vitro electrophoretic mobility shift assay (EMSA). This whitepaper proposes a synergistic, complementary workflow that leverages the unique strengths of both techniques for robust and conclusive target validation in drug discovery pipelines.
The Complementary Paradigm
ChIP-seq provides a genome-wide, in vivo snapshot of TF binding events within their native chromatin context, identifying potential regulatory regions. However, it can yield false positives due to indirect binding or technical artifacts. EMSA offers direct, in vitro biochemical verification of protein-nucleic acid interaction with precise control over reaction components, but lacks genomic scale and cellular context. The integrated workflow uses ChIP-seq for discovery and EMSA for rigorous validation of specific interactions.
Quantitative Comparison of Core Methodologies
Table 1: Comparative Analysis of ChIP-seq and EMSA
| Feature | ChIP-seq | EMSA (Classical) |
|---|---|---|
| Binding Context | In vivo (native chromatin) | In vitro (purified components) |
| Throughput | Genome-wide (high) | Single locus/probe (low) |
| Primary Output | Binding regions (peaks) | Confirmation of direct binding |
| Key Quantitative Metric | Peak enrichment (FDR q-value, fold-change) | Shifted probe intensity (% bound) |
| Typical Resolution | 100-1000 bp | Exact binding site (20-40 bp oligo) |
| Time to Result | 3-5 days (post-library prep) | 1-2 days |
| Critical Reagent | High-quality, specific antibody | Purified TF protein / nuclear extract |
Proposed Complementary Workflow
The following diagram illustrates the sequential, iterative workflow for target validation.
Diagram Title: Complementary ChIP-seq & EMSA Workflow for TF Target Validation
Detailed Experimental Protocols
Protocol 1: ChIP-seq for TF Binding Site Discovery
- Crosslinking & Cell Lysis: Treat cells (e.g., 10^7) with 1% formaldehyde for 10 min at RT. Quench with 125mM glycine. Harvest and lyse in SDS lysis buffer.
- Chromatin Shearing: Sonicate lysate to fragment DNA to 200-500 bp. Confirm size by agarose gel electrophoresis.
- Immunoprecipitation: Incubate chromatin with 2-5 µg of validated, high-specificity anti-TF antibody overnight at 4°C. Use Protein A/G beads for capture.
- Washing & Elution: Wash beads sequentially with Low Salt, High Salt, LiCl, and TE buffers. Elute complexes with freshly prepared elution buffer (1% SDS, 0.1M NaHCO3).
- Reverse Crosslinks & Purification: Incubate eluate with 200mM NaCl at 65°C overnight. Treat with RNase A and Proteinase K. Purify DNA using silica-membrane columns.
- Library Prep & Sequencing: Prepare sequencing library from ChIP and Input control DNA using a commercial kit (e.g., NEBNext Ultra II). Sequence on an Illumina platform (≥20 million reads/sample).
Protocol 2: EMSA forIn VitroBinding Validation
- Protein Preparation: Use purified recombinant TF protein or prepare nuclear extract from relevant cell lines.
- Probe Labeling: End-label 20-30 bp double-stranded oligonucleotide containing the candidate ChIP-seq peak sequence with [γ-32P] ATP using T4 Polynucleotide Kinase. Purify using a spin column.
- Binding Reaction: Assemble 20 µL reaction: 4 µL 5X binding buffer (50mM Tris, 250mM NaCl, 5mM DTT, 30% glycerol, 5mM MgCl2), 2 µg poly(dI-dC), 1-10 µg nuclear extract or 10-100 ng purified TF, 10 fmol labeled probe. Incubate 20 min at RT.
- Electrophoresis: Load reaction onto a pre-run 6% non-denaturing polyacrylamide gel in 0.5X TBE buffer. Run at 100V for 60-90 min at 4°C.
- Detection: Transfer gel to filter paper, dry, and expose to a phosphorimager screen. Quantify band intensities.
- Specificity Controls: Include reactions with a 100x molar excess of unlabeled cold competitor (specific or mutant) and an antibody for supershift confirmation.
Pathway Visualization of Validation Logic
The logical relationship between experimental outcomes and conclusions is critical.
Diagram Title: EMSA Validation Controls & Interpretations
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for the Complementary Workflow
| Reagent Category | Specific Item | Function & Critical Consideration |
|---|---|---|
| Antibodies | Validated ChIP-seq Grade Anti-TF Antibody | Immunoprecipitates the target TF for ChIP-seq. Specificity is paramount; knock-out/knockdown validation preferred. |
| Control IgG (Species-matched) | Negative control for non-specific IP in ChIP-seq. | |
| Assay Kits | Commercial ChIP-seq Kit (e.g., Cell Signaling, Abcam) | Provides optimized buffers, beads, and protocols for robust, reproducible ChIP. |
| Radiolabeling Kit (e.g., T4 PNK) | For efficient end-labeling of EMSA probes with 32P. Non-radioactive alternatives (chemiluminescent) exist. | |
| Molecular Biology | NEBNext Ultra II DNA Library Prep Kit | High-efficiency library preparation from low-input ChIP DNA for sequencing. |
| Poly(dI-dC) | Non-specific competitor DNA in EMSA to reduce non-specific protein-probe binding. | |
| Probes & Oligos | Biotin- or 32P-labeled Double-stranded Oligonucleotides | EMSA probes representing top ChIP-seq peaks and mutated controls for binding specificity. |
| Protein Tools | Recombinant Purified TF Protein (full-length or DBD) | Gold standard for EMSA, ensuring the observed shift is due to the TF alone. |
| Nuclear Extract Kit (e.g., from NE-PER) | Source of native TF and co-factors for more physiologically relevant EMSA. |
This complementary workflow transforms the perceived dichotomy between ChIP-seq and EMSA into a powerful, iterative engine for target validation. By employing ChIP-seq as a discovery platform and EMSA as a definitive biochemical filter, researchers can build a highly confident shortlist of direct TF-target interactions. This rigorous, two-tiered approach de-risks downstream functional assays and provides a solid foundation for drug discovery programs aimed at modulating transcriptional networks.
Solving Common Problems and Enhancing Assay Performance
Within the framework of transcription factor (TF) binding research, the Electrophoretic Mobility Shift Assay (EMSA) serves as a foundational in vitro technique for validating direct protein-nucleic acid interactions. While high-throughput methods like ChIP-seq provide genome-wide binding maps in vivo, EMSA offers indispensable mechanistic proof of direct binding and allows for precise biochemical characterization of binding affinity and specificity. This guide addresses the core technical challenges—non-specific binding, smearing, and weak shifts—that can obscure data interpretation and compromise the bridge between in silico prediction, in vitro validation, and in vivo relevance.
Table 1: Prevalence and Impact of Common EMSA Artifacts
| Artifact | Reported Frequency in Problematic Assays | Primary Consequence | Typical Success Rate Post-Optimization |
|---|---|---|---|
| Non-Specific Binding | ~65% | False positives, obscured specific shift | >90% |
| Smearing | ~50% | Uninterpretable bands, poor resolution | >85% |
| Weak or No Shift | ~45% | False negatives, failed validation | 70-80% |
Table 2: Optimized Component Concentrations for Troubleshooting
| Reaction Component | Standard Range | For Non-Specific Binding | For Smearing | For Weak Shift |
|---|---|---|---|---|
| Poly(dI:dC) | 0.05-0.1 µg/µL | 0.1-0.5 µg/µL | 0.05-0.1 µg/µL | 0.01-0.05 µg/µL |
| NP-40/Tween-20 | 0-0.1% | 0.1-0.5% | 0% | 0-0.1% |
| Glycerol | 0-2.5% | 2.5% | <1% | 2.5% |
| NaCl/KCl | 50-100 mM | >150 mM | 50 mM | <50 mM |
| MgCl₂ | 0-10 mM | 5 mM | 0 mM | 5-10 mM |
| Probe (Labeled) | 0.1-1 nM | 0.1 nM | 0.1 nM | 1-2 nM |
| Protein (Lysate) | 2-10 µg | 2 µg | 2-5 µg | 5-20 µg |
Detailed Experimental Protocols
Protocol 1: Systematic Optimization to Minimize Non-Specific Binding
- Probe Design & Preparation: Use HPLC-purified oligonucleotides. Label probe with γ-³²P ATP or a fluorescent dye via end-labeling. Gel-purify the labeled probe.
- Competitor Titration: In a 20 µL binding reaction (10 mM HEPES pH 7.9, 50-150 mM KCl, 1 mM DTT, 2.5% glycerol, 0.1 mM EDTA), include constant amounts of nuclear extract (5 µg) and labeled probe (0.1 nM, 20,000 cpm). Set up a parallel series with increasing poly(dI:dC) (0, 0.05, 0.1, 0.25, 0.5 µg/µL final). Incubate 20 min at RT.
- Detergent Optimization: Add non-ionic detergents (NP-40 or Tween-20 at 0.1%, 0.25%, 0.5%) to reactions with optimized poly(dI:dC).
- Electrophoresis: Pre-run a 6% non-denaturing polyacrylamide gel (29:1 acrylamide:bis) in 0.5x TBE at 100V for 30 min at 4°C. Load samples, run at 100V for 60-90 min with circulation of cold buffer.
- Analysis: Expose gel to phosphorimager or X-ray film. The optimal condition yields a sharp, discrete shifted band with minimal background in the probe lane.
Protocol 2: Eliminating Smearing and Improving Band Resolution
- Reaction Cleanliness: Ensure all buffers are filtered (0.22 µm) and degassed. Use high-purity, nuclease-free water.
- Salt & Divalent Cation Adjustment: Prepare binding reactions with lower ionic strength (50 mM KCl) and omit MgCl₂ initially. Titrate MgCl₂ back (1, 2, 5 mM) if the shift is Mg²⁺-dependent.
- Glycerol Reduction: Reduce glycerol concentration to ≤1% to prevent complex aggregation.
- Gel Electrophysis Parameters: Increase gel percentage to 8% for larger complexes. Ensure the gel apparatus is thoroughly cleaned. Run gel at 4°C with buffer recirculation. Load samples without bromophenol blue (which can interfere), using a separate dye lane if needed.
- Post-Electrophoresis Handling: Carefully transfer gel onto Whatman paper, dry under vacuum at 80°C for 1 hour before autoradiography to prevent diffusion.
Protocol 3: Enhancing Weak or Absent Shifts
- Protein Source Verification: Use fresh nuclear extract or purified protein. Confirm protein activity via a positive control DNA probe.
- Probe Excess: Increase the amount of labeled probe to 1-2 nM while keeping protein constant.
- Protein Concentration & Incubation: Titrate protein amount upward (10-20 µg). Extend incubation time to 30 min at 4°C (instead of RT) to promote complex stability.
- Cofactor Addition: Add essential cofactors identified in the literature (e.g., Zn²⁺ for zinc-finger TFs, specific lipids).
- Antibody Supershift: If a specific antibody is available, add 1-2 µg to the reaction after initial binding and incubate further 20 min. This can stabilize the complex and cause a further mobility shift ("supershift").
Visualizing EMSA Workflow and Troubleshooting Logic
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Robust EMSA
| Reagent/Material | Function & Rationale | Recommended Product Example (Illustrative) |
|---|---|---|
| Poly(dI:dC) | Non-specific competitor DNA; absorbs non-sequence-specific DNA-binding proteins to reduce background. | Sigma-Aldrich P4929 |
| Non-Ionic Detergent (NP-40/Tween-20) | Reduces hydrophobic protein-protein aggregation and non-specific binding. | Thermo Fisher Scientific 28324 (NP-40) |
| γ-³²P ATP | Radioactive label for high-sensitivity probe detection via autoradiography. | PerkinElmer NEG002Z |
| Chemiluminescent Nucleic Acid Labeling Kit | Non-radioactive alternative for biotin or digoxigenin labeling and detection. | Thermo Fisher Scientific 20160 (LightShift Chemiluminescent EMSA Kit) |
| High-Purity Acrylamide/Bis (29:1) | For casting reproducible, non-denaturing gels with consistent pore size. | Bio-Rad 1610156 |
| Protease & Phosphatase Inhibitor Cocktails | Preserves TF integrity and phosphorylation state in crude extracts. | Roche cOmplete, PhosSTOP |
| Non-Specific Competitor (e.g., salmon sperm DNA) | Alternative to poly(dI:dC) for some TFs; requires titration. | Invitrogen 15632011 |
| Mobility Shift Buffer (10X) | Consistent buffer formulation (HEPES, KCl, DTT, glycerol) for binding reactions. | Thermo Fisher Scientific 20158 |
| TF-Specific Antibody | For supershift experiments to confirm TF identity in complex. | Vendor and clone specific to TF under study. |
| Cooled Circulating Electrophoresis Unit | Maintains 4°C during run to prevent complex dissociation and smearing. | Bio-Rad Model 1000 Chiller |
1. Introduction: Within the ChIP-seq vs. EMSA Paradigm
In transcription factor (TF) binding research, the choice between Chromatin Immunoprecipitation followed by sequencing (ChIP-seq) and the Electrophoretic Mobility Shift Assay (EMSA) hinges on the trade-off between in vivo genomic context and in vitro biochemical specificity. While EMSA offers precise, quantitative analysis of protein-DNA interactions under controlled conditions, it lacks the genomic-scale, in vivo resolution of ChIP-seq. The power of ChIP-seq to map TF binding sites across the entire genome is, however, critically dependent on three technical pillars: antibody specificity, chromatin fragmentation efficiency, and optimal signal-to-noise ratio. This guide provides an in-depth troubleshooting framework for these core challenges, ensuring data quality that robustly validates in vivo findings suggested by in vitro assays like EMSA.
2. Pillar I: Validating and Improving Antibody Specificity
Antibody specificity is the foremost determinant of ChIP-seq success. Non-specific antibodies generate high background noise, obscuring true binding signals.
Experimental Protocol: Antibody Validation Pre-ChIP
- Knockdown/Knockout Validation: Perform siRNA-mediated knockdown or CRISPR-Cas9 knockout of the target TF in your cell line. Conduct a standard ChIP-qPCR on known positive and negative genomic regions. A specific antibody will show >70% signal reduction at positive sites in knockdown/knockout samples compared to wild-type controls.
- Peptide Blocking Competition: Pre-incubate the ChIP antibody with a 5-10 fold molar excess of the immunogenic peptide (or a recombinant protein epitope) for 1 hour at 4°C before adding to the chromatin. The blocked antibody should show >80% reduction in ChIP-qPCR signal compared to the untreated antibody control.
- Western Blot Analysis: Use the ChIP antibody for western blot on whole-cell and nuclear extracts. A specific antibody should produce a single dominant band at the expected molecular weight, with minimal non-specific bands.
Table 1: Antibody Validation Strategies & Success Criteria
| Validation Method | Experimental Readout | Quantitative Success Criteria |
|---|---|---|
| Genetic Knockdown/KO | ChIP-qPCR at known sites | >70% signal reduction |
| Peptide Blocking | ChIP-qPCR at known sites | >80% signal reduction |
| Western Blot | Banding pattern on lysates | Single dominant band at correct MW |
3. Pillar II: Optimizing Sonication for Efficient Chromatin Fragmentation
Uniform chromatin shearing to 100-500 bp fragments is essential for resolution and library compatibility. Under-sonication reduces resolution; over-sonication damages epitopes.
Experimental Protocol: Sonication Optimization
- Cell Lysis: Crosslink 1-10 x 10^6 cells with 1% formaldehyde for 10 min. Quench with 125 mM glycine. Pellet cells, wash with cold PBS. Resuspend in LB1 buffer (50 mM HEPES-KOH pH 7.5, 140 mM NaCl, 1 mM EDTA, 10% Glycerol, 0.5% NP-40, 0.25% Triton X-100) for 10 min at 4°C. Pellet, resuspend in LB2 buffer (10 mM Tris-HCl pH 8.0, 200 mM NaCl, 1 mM EDTA, 0.5 mM EGTA) for 10 min at 4°C.
- Sonication Test: Resuspend pellet in 1 mL of Shearing Buffer (0.1% SDS, 1 mM EDTA, 10 mM Tris-HCl pH 8.0). Split into 6 identical 150 µL aliquots in 1.5 mL tubes. Using a focused ultrasonicator (e.g., Covaris, Bioruptor), subject each aliquot to a different number of cycles (e.g., 2, 4, 6, 8, 10, 12 cycles; 30 sec ON/30 sec OFF per cycle, 4°C).
- Analysis: Reverse crosslinks for all aliquots at 65°C overnight with 200 mM NaCl. Treat with RNase A and Proteinase K. Purify DNA. Analyze 20 µL on a 2% agarose gel or a Bioanalyzer/TapeStation. The optimal condition yields a smear centered at 200-300 bp.
Table 2: Troubleshooting Sonication Efficiency
| Problem | Possible Cause | Solution |
|---|---|---|
| Large fragments (>1000 bp) | Under-sonication, low cell number | Increase cycles/duration; ensure correct cell count |
| Over-fragmentation (<150 bp) | Over-sonication, high power | Reduce cycles/duration; decrease power setting |
| Inconsistent shearing | Variable sample volume, foaming | Use identical, flat-cap tubes; avoid air bubbles |
Title: Sonication Optimization Workflow
4. Pillar III: Enhancing Signal-to-Noise Ratio (SNR)
High SNR is critical for distinguishing true peaks from background. Key factors include specific vs. non-specific DNA purification and sequencing depth.
Experimental Protocol: Using Spike-in Controls for Normalization
- Spike-in Chromatin Addition: Use chromatin from a different species (e.g., Drosophila melanogaster S2 cells) as a spike-in. Add a fixed amount (e.g., 1-10% of experimental chromatin) to your human/mouse samples after sonication but before immunoprecipitation.
- Immunoprecipitation: Proceed with your target antibody. The antibody should not recognize spike-in chromatin, so its recovery is proportional to non-specific background.
- qPCR & Analysis: Design qPCR primers for your target's known binding sites (experimental) and for the spike-in genome (background). Calculate the % input for both. A high SNR is indicated by a high experimental % input relative to the spike-in % input. For sequencing, use spike-in aligned reads to normalize between samples quantitatively.
Table 3: Strategies to Improve Signal-to-Noise Ratio
| Strategy | Mechanism | Expected Outcome |
|---|---|---|
| Spike-in Normalization | Controls for technical variation in IP efficiency | Enables accurate cross-sample comparison |
| Increased Sequencing Depth | Better statistical power for peak calling | Identifies lower-affinity binding sites |
| Stringent Washes | Reduces non-specific antibody binding | Lowers background, sharpens peaks |
| Control Experiments (IgG, Input) | Defines background model for peak callers | Reduces false positive rate |
5. The Scientist's Toolkit: Essential Research Reagent Solutions
Table 4: Key Reagents for Robust ChIP-seq
| Reagent/Material | Function & Importance | Example/Typical Use |
|---|---|---|
| Validated ChIP-grade Antibody | Specifically enriches target protein-DNA complexes; the single most critical reagent. | Antibodies with published ChIP-seq datasets (e.g., from Abcam, Cell Signaling, Diagenode). |
| Magnetic Protein A/G Beads | Efficient capture of antibody complexes with low non-specific binding. | Dynabeads, Protein A/G Magnetic Beads. |
| Crosslinking Reagent (Formaldehyde) | Reversible fixation of protein-DNA interactions in vivo. | 1% final concentration, 10 min incubation. |
| Sonication Device | Fragments chromatin to optimal size for high-resolution mapping. | Focused ultrasonicator (Covaris), bath sonicator (Bioruptor). |
| Spike-in Chromatin | Exogenous chromatin for normalization, correcting for technical variation. | Drosophila S2 chromatin (e.g., from Active Motif). |
| DNA Clean-up Beads/Columns | Purify immunoprecipitated DNA with high recovery for low-input libraries. | SPRIselect beads, MinElute PCR Purification columns. |
| High-Sensitivity DNA Assay | Accurate quantification of picogram amounts of ChIP DNA pre-library prep. | Qubit dsDNA HS Assay, TapeStation High Sensitivity D1000. |
| Library Prep Kit for Low Input | Converts low-mass ChIP DNA into sequencing libraries with minimal bias. | KAPA HyperPrep, NEBNext Ultra II DNA Library Prep. |
Title: Signal-to-Noise Optimization Workflow
6. Conclusion: Integrating Robust ChIP-seq with EMSA
A meticulously optimized ChIP-seq protocol, with validated antibodies, efficient sonication, and controlled SNR, produces high-confidence in vivo binding maps. These maps provide the essential genomic context to frame and interpret the precise, quantitative protein-DNA affinities measured by in vitro EMSA. Together, they form a complementary and powerful pipeline for definitive transcription factor research, bridging biochemical mechanism with cellular function.
Within the broader methodology for studying transcription factor (TF)-DNA interactions, Electrophoretic Mobility Shift Assay (EMSA) remains a foundational, in vitro technique. It is often contrasted with in vivo methods like Chromatin Immunoprecipitation followed by sequencing (ChIP-seq). While ChIP-seq maps genome-wide binding events within a cellular context, EMSA provides biochemical validation of direct, sequence-specific binding, offering definitive proof of interaction that ChIP-seq alone cannot. This guide details advanced optimization strategies for EMSA—cold competitors, buffer conditions, and supershifts—to generate robust, publication-quality data that can effectively complement and validate ChIP-seq findings.
Core Principles & Optimization Strategies
The Role of Cold Competitor Probes
Cold competitor oligonucleotides are identical, unlabeled DNA probes used to demonstrate binding specificity. Their effectiveness is concentration-dependent.
Table 1: Cold Competitor Optimization Data
| Competitor Type | Typical Molar Excess (vs. labeled probe) | Expected Outcome | Interpretation |
|---|---|---|---|
| Specific (Unlabeled) | 10x - 100x | Complete abolition of shifted band | Confirms sequence-specific binding. |
| Non-specific (e.g., poly(dI:dC)) | 50x - 200x | No reduction in shifted band | Confirms lack of non-specific interference. |
| Mutant (cis-element mutated) | 100x | No or minimal competition | Confirms exact sequence requirement. |
Protocol: Cold Competitor Titration
- Prepare Competitor Mixes: Set up a series of binding reactions with a constant amount of nuclear extract (e.g., 5 µg) and labeled probe (e.g., 20 fmol).
- Add Competitor: Include increasing molar excesses (0x, 10x, 25x, 50x, 100x) of unlabeled specific competitor or non-specific DNA (e.g., poly(dI:dC)).
- Incubate: Pre-incubate extract with competitor on ice for 10 minutes before adding the hot probe. This allows competitors to bind first.
- Proceed: Add labeled probe, incubate 20-30 min at room temperature, and run on a native gel.
- Analyze: Quantify band intensity. Specific binding should be progressively abolished with specific, but not non-specific, competitors.
Critical Buffer Conditions
Buffer composition dictates complex stability and specificity.
Table 2: Key Buffer Components and Optimization Ranges
| Component | Typical Concentration Range | Function | Optimization Effect |
|---|---|---|---|
| Buffer (pH) | 10 mM HEPES (pH 7.5-7.9) | Maintains pH | Affects protein folding and binding affinity. |
| KCl/NaCl | 50-150 mM | Controls ionic strength | High salt (>200 mM) disrupts electrostatic interactions; low salt may increase non-specific binding. |
| MgCl₂ | 1-5 mM | Divalent cation | Often stabilizes protein-DNA complexes; test with/without. |
| DTT/β-Mercaptoethanol | 1 mM DTT | Reducing agent | Maintains cysteine residues in reduced state; critical for some TFs. |
| Non-ionic Detergent (NP-40/Triton X-100) | 0.1% | Reduces non-specific binding | Minimizes protein adherence to tubes. |
| Carrier Protein (BSA) | 0.1 mg/mL | Stabilizes proteins | Reduces non-specific loss; not always required. |
| Glycerol | 5-10% | Increases viscosity | Aids in loading; can stabilize some complexes. |
| Poly(dI:dC) | 0.05-0.1 mg/mL | Non-specific DNA competitor | Blocks non-specific protein-DNA interactions. Titrate carefully. |
Protocol: Buffer Condition Screening
- Prepare Master Stock Solutions: Create 5X stock solutions of buffers varying in salt (KCl: 50, 100, 150 mM), MgCl₂ (0, 2, 5 mM), and non-ionic detergent (0, 0.05%).
- Set Up Reaction Matrix: In a 96-well plate or PCR strips, assemble binding reactions where each condition combines different buffer components.
- Run EMSA: Keep protein, probe, and incubation time constant across all conditions.
- Evaluate: Assess which condition yields the sharpest, most intense specific complex with minimal non-specific smearing.
Supershifts for Complex Identification
A supershift occurs when an antibody against the bound protein further retards the complex, confirming TF identity.
Protocol: Supershift Assay
- Form Complex: Perform standard binding reaction with protein extract and labeled probe. Incubate for 20 min.
- Add Antibody: Add 1-2 µg of specific antibody (or corresponding IgG control). Do not add detergent (SDS) containing buffers.
- Secondary Incubation: Incubate further for 30-60 minutes on ice or at 4°C.
- Load and Run: Load entire reaction on a pre-run, pre-cooled native gel. Run at lower voltage (e.g., 100V) for longer to resolve the heavier supershifted complex.
- Controls: Always include: a) Probe alone, b) Protein + probe, c) Protein + probe + specific antibody, d) Protein + probe + non-specific/isotype control antibody.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Optimized EMSA
| Item | Function & Selection Criteria |
|---|---|
| Chemiluminescent EMSA Kit | Non-radioactive detection (e.g., biotin- or digoxigenin-labeled probes). Includes labeling, binding, and detection reagents. |
| HEK 293T Nuclear Extract | Common positive control source for many human TFs. Ensures experiment functionality. |
| Recombinant Transcription Factor | Pure protein for establishing baseline binding without confounding factors from crude extracts. |
| Gel Shift Binding 5X Buffer | Commercial optimized buffer (e.g., from Thermo Fisher). Good starting point for difficult TFs. |
| Poly(dI:dC) | Synthetic non-specific DNA competitor. Critical for reducing non-specific shifts in crude extracts. |
| Non-radioactive Probe Labeling Kit | For safe, stable labeling of oligonucleotides with biotin or DIG. |
| High-Density TBE Buffer (5X) | For preparing native polyacrylamide gels. Ensures consistent pH and conductivity. |
| Pre-cast Native PAGE Gels (6%) | Ensure consistency and save time in gel preparation. |
| TF-Specific Antibody (ChIP-grade) | High-affinity, well-validated antibody for supershift assays. Must recognize native protein. |
| Magnetic Shift EMSA Kit | Solution-based EMSA using streptavidin-magnetic beads and colorimetric/chemiluminescent readout. Avoids gel electrophoresis. |
Visualizing Workflows and Relationships
Title: EMSA Optimization & Validation Workflow
Title: Complementary Strengths of ChIP-seq and EMSA
A meticulously optimized EMSA, employing titrated cold competitors, refined buffer conditions, and conclusive supershifts, provides an indispensable layer of biochemical rigor to transcription factor research. When integrated with the genomic landscape revealed by ChIP-seq, it forges a powerful complementary approach. This combination moves from observing where a factor binds in the genome to definitively proving how it interacts with a specific DNA sequence, a critical step for understanding gene regulation and validating therapeutic targets.
This technical guide addresses three critical, interdependent parameters for a successful Chromatin Immunoprecipitation followed by sequencing (ChIP-seq) experiment: crosslinking time, sequencing depth, and peak calling settings. The optimization of this triad is framed within a broader thesis comparing ChIP-seq to the Electrophoretic Mobility Shift Assay (EMSA) for transcription factor (TF) binding research. While EMSA offers in vitro specificity for protein-nucleic acid interactions, ChIP-seq provides a genome-wide, in vivo snapshot of binding events within their native chromatin context. The choice between these techniques hinges on the research question: EMSA for mechanistic, reductionist validation of direct binding, and ChIP-seq for discovering the genomic landscape of TF occupancy. This guide focuses on refining ChIP-seq to yield data of a quality that allows for robust biological inference, minimizing false positives and negatives.
Core Parameter 1: Crosslinking Time Optimization
Crosslinking captures transient protein-DNA interactions. Insufficient crosslinking leads to poor yield; excessive crosslinking causes epitope masking, chromatin fragmentation issues, and increased background noise.
Experimental Protocol: Crosslinking Time Course
Objective: To determine the optimal formaldehyde crosslinking time for a specific transcription factor-cell type combination. Materials: Adherent or suspension cells, 37% formaldehyde, 2.5M glycine, PBS, cell scraper, conical tubes. Procedure:
- Grow cells to ~80% confluence.
- For each time point (e.g., 1, 5, 10, 15, 30 min), directly add 37% formaldehyde to culture medium to a final concentration of 1%.
- Incubate at room temperature with gentle agitation.
- Quench crosslinking by adding 2.5M glycine to a final concentration of 0.125M. Incubate for 5 min.
- Wash cells twice with cold PBS.
- Harvest cells by scraping (adherent) or centrifugation. Pellet can be flash-frozen.
- Perform parallel ChIP-seq experiments for the target TF and an IgG control for each time point.
- Assess yield via qPCR at positive control genomic loci and signal-to-noise ratio.
Table 1: Impact of Crosslinking Time on ChIP-seq Outcomes
| Crosslinking Time (min) | DNA Yield | Fragmentation Efficiency | Signal-to-Noise Ratio | Risk of Epitope Masking |
|---|---|---|---|---|
| 1-2 | Low | High | Variable, often low | Very Low |
| 5-10 | Optimal for most TFs | High | High | Low |
| 15-20 | High | Moderate | High | Moderate |
| ≥30 | Very High | Poor (difficult to sonicate) | Low (high background) | High |
Core Parameter 2: Sequencing Depth Determination
Sequencing depth (total number of reads) directly impacts the sensitivity and reproducibility of peak detection, especially for low-abundance TFs or broad histone marks.
Experimental Protocol: Saturation Analysis
Objective: To determine the minimum read depth required for confident peak calling. Materials: High-quality ChIP-seq library, sequencing facility access. Procedure:
- Sequence one library to a very high depth (e.g., 80-100 million reads for a mammalian genome).
- Randomly subsample the full dataset (e.g., 10%, 20%, 30%...90% of reads) using bioinformatics tools like
seqtkor SAMtools. - Perform peak calling on each subsampled dataset with fixed parameters.
- For each depth, calculate the proportion of peaks identified relative to the full dataset and plot a saturation curve. The point where the curve plateaus indicates sufficient depth.
Table 2: Recommended Sequencing Depth Guidelines (Mammalian Genomes)
| Target Type | Example | Minimum Recommended Depth | Optimal Depth | Rationale |
|---|---|---|---|---|
| Point-source TF | p53, CTCF | 10-15 million reads | 20-30 million reads | Sharp, narrow peaks; moderate depth yields high confidence. |
| Pioneer Factor / Broad TF | FOXA1, Pol II | 20-30 million reads | 40-60 million reads | Broader enrichment regions require more reads for full coverage. |
| Histone Mark (Promoter) | H3K4me3 | 15-20 million reads | 25-40 million reads | Sharp, defined peaks at promoters. |
| Histone Mark (Enhancer/ Broad) | H3K27me3, H3K36me3 | 30-40 million reads | 50-80 million reads | Very broad domains require extensive sampling. |
Core Parameter 3: Peak Calling Parameter Tuning
Peak calling algorithms (e.g., MACS2, HOMER) use statistical models to distinguish true enrichment from background. Key parameters include the p-value/q-value threshold and the shift size.
Experimental Protocol: Parameter Grid Evaluation
Objective: To optimize peak caller settings for a specific experiment. Materials: Aligned ChIP-seq and input control BAM files, peak calling software (e.g., MACS2). Procedure:
- Run MACS2
callpeakwith a grid of parameters:-p(p-value): Test 1e-3, 1e-5, 1e-7.-q(q-value/FDR): Test 0.01, 0.05, 0.10.--shift/--extsize: Test predicted fragment length from cross-correlation analysis.
- For each resulting peak set, evaluate quality metrics:
- FRiP (Fraction of Reads in Peaks): A higher FRiP (>1% for TFs, >10% for histone marks) indicates a successful experiment.
- Irreproducible Discovery Rate (IDR): Compare replicates to assess consistency. Peaks with IDR < 0.05 are highly reproducible.
- Visual Inspection (IGV): Validate top peaks and borderline calls.
- Select parameters that maximize high-confidence, reproducible peaks.
Table 3: Effect of Peak Calling Stringency on Output
| Parameter Setting | Number of Peaks Called | False Discovery Rate (FDR) | Stringency | Recommended Use Case |
|---|---|---|---|---|
| p-value=1e-3, q-value=0.1 | Very High | High (>10%) | Low | Exploratory analysis, initial scan. |
| p-value=1e-5, q-value=0.05 | High | Moderate (5%) | Moderate | Standard balance for most TFs. |
| p-value=1e-7, q-value=0.01 | Moderate | Low (<1%) | High | Conservative list for validation (e.g., EMSA follow-up). |
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Materials for Optimized ChIP-seq
| Item | Function | Example/Consideration |
|---|---|---|
| High-Quality Antibody | Specifically immunoprecipitates the target antigen. | Validate for ChIP-seq suitability (ChIP-grade). High specificity is non-negotiable. |
| Ultrapure Formaldehyde | Crosslinks proteins to DNA and proteins to proteins. | Use fresh, high-purity (e.g., methanol-free) for consistent efficiency. |
| Magnetic Protein A/G Beads | Capture antibody-target complexes. | Superior recovery and lower background vs. agarose/salmon sperm slurries. |
| Covaris/Sonicator | Shears crosslinked chromatin to optimal fragment size (200-600 bp). | Adaptive Focused Acoustics (Covaris) provides most consistent shear profile. |
| SPRI Beads (e.g., AMPure) | Size selection and clean-up of DNA libraries. | Efficient, automatable replacement for gel electrophoresis. |
| Library Prep Kit | Prepares sequencing library from immunoprecipitated DNA. | Kits with low input and UMI support are advantageous. |
| High-Sensitivity DNA Assay | Quantifies low-concentration DNA (ChIP DNA, libraries). | Critical for accurate library pooling (e.g., Qubit, Bioanalyzer). |
Visualizing the Optimization Workflow and Thesis Context
Diagram Title: ChIP-seq Optimization Workflow within the ChIP-seq vs. EMSA Thesis
Diagram Title: Impact of Parameter Choices on ChIP-seq Experimental Outcomes
Within the context of transcription factor (TF) binding research, Chromatin Immunoprecipitation followed by sequencing (ChIP-seq) and Electrophoretic Mobility Shift Assay (EMSA) are cornerstone techniques. ChIP-seq maps in vivo genome-wide binding sites, while EMSA probes in vitro protein-nucleic acid interactions with precise biochemical resolution. A major challenge in the field is the reproducibility and specificity of data generated by these methods. This guide details the critical, often parallel, controls required for both assays to validate findings and ensure robust, interpretable results.
Core Control Strategies: A Comparative Framework
The following table summarizes the essential controls for ChIP-seq and EMSA, highlighting their shared principles and assay-specific implementations.
Table 1: Critical Controls for ChIP-seq and EMSA
| Control Objective | ChIP-seq Implementation | EMSA Implementation | Purpose & Rationale |
|---|---|---|---|
| Specificity of TF-Probe Interaction | Use of isotype control IgG; Knockout/knockdown cell lines; Competition with free cognate oligonucleotide. | Competition with unlabeled (cold) specific probe; Use of mutant/non-specific cold probe. | Distinguishes specific binding from non-specific antibody interactions (ChIP) or protein-probe interactions (EMSA). |
| Antibody Validation | Primary antibody specificity verified by siRNA/shRNA, knockout, or orthogonal assay (e.g., EMSA). | Antibody for supershift: Must be validated for target epitope accessibility and non-interference. | Ensures the antibody recognizes the target TF and does not produce spurious signals. |
| Input DNA/Nuclear Extract Quality | Sequencing of Input DNA (non-immunoprecipitated chromatin). | Verification of nuclear extract activity via a positive control probe/protein combination. | Controls for chromatin accessibility/sequencing bias (ChIP-seq) and confirms extract functionality (EMSA). |
| Background & Non-Specific Binding | No-antibody control (beads-only); Use of genomic regions known to lack binding (negative genomic loci). | Incubation with non-specific competitor (e.g., poly(dI-dC)); Probe-only lane. | Measures and minimizes background from bead capture or non-specific protein-DNA interactions. |
| Reproducibility | Biological replicates (≥2); High correlation of peak calls (IDR analysis). | Technical replicates of binding reactions; Quantification from multiple gel images. | Assesses experimental variability and statistical confidence of identified binding events. |
| Signal Normalization | Spike-in controls (e.g., Drosophila chromatin, exogenous cells). | Use of a constitutively binding complex or labeled control probe. | Corrects for technical variation in IP efficiency or loading/transfer differences (EMSA). |
| Functional Validation | Motif enrichment analysis within peaks; Correlation with gene expression (RNA-seq). | Mutation of consensus binding site leading to loss of shift; Supershift with a second, independent antibody. | Confirms the biological relevance and sequence specificity of the observed interaction. |
Detailed Experimental Protocols for Key Controls
ChIP-seq: Isotype Control and Input DNA Preparation
Protocol:
- Chromatin Preparation: Cross-link cells with 1% formaldehyde for 10 min at room temperature. Quench with 125 mM glycine. Sonicate chromatin to an average fragment size of 200-500 bp.
- Immunoprecipitation Split: Divide sheared chromatin into three aliquots:
- Test IP: Incubate with target TF-specific antibody.
- Isotype Control IP: Incubate with a non-specific antibody of the same isotype (e.g., normal rabbit IgG) at the same concentration.
- Input Control: Set aside 10% of sheared chromatin, reverse cross-links, and purify DNA.
- Common Steps: All IPs use protein A/G magnetic beads, followed by sequential washes (Low Salt, High Salt, LiCl, TE buffers). Reverse cross-links at 65°C overnight.
- DNA Purification: Treat with RNase A and Proteinase K, then purify DNA using silica-membrane columns.
- Library Preparation & Sequencing: Prepare sequencing libraries from Test IP, Isotype Control, and Input DNA. Sequence on an appropriate platform (e.g., Illumina). The Input DNA serves as the control for genomic sequencing bias.
EMSA: Specific and Non-Specific Competition Assay
Protocol:
- Probe Labeling: Label 20-50 fmol of double-stranded oligonucleotide containing the putative binding site with [γ-³²P]ATP using T4 Polynucleotide Kinase. Purify using a spin column.
- Binding Reaction Setup: Prepare reactions in a 20 µL volume containing:
- 1X Binding Buffer (10 mM HEPES, pH 7.9, 50 mM KCl, 1 mM DTT, 2.5% glycerol, 0.05% NP-40).
- 2 µg poly(dI-dC) as non-specific competitor.
- 5-10 µg nuclear extract or purified TF protein.
- Competitor Conditions: a. No competitor: Add water. b. Specific competitor: Add 50x or 100x molar excess of unlabeled identical probe. c. Non-specific competitor: Add 50x or 100x molar excess of unlabeled probe with a scrambled/mutated sequence.
- Incubation: Pre-incubate extract with competitors on ice for 10 min. Add labeled probe and incubate at room temperature for 20 min.
- Electrophoresis: Load samples onto a pre-run, non-denaturing 5-6% polyacrylamide gel in 0.5X TBE buffer. Run at 100-150 V at 4°C until the free probe migrates near the bottom.
- Detection: Dry gel and expose to a phosphorimager screen. Specific binding is indicated by a shifted band that is abolished by excess specific cold probe, but not by excess non-specific cold probe.
Visualization of Control Strategies
Control Strategy Decision Flow for ChIP-seq & EMSA
EMSA Competition Assay Logic
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Critical Controls
| Reagent / Material | Primary Assay | Function in Control Experiments |
|---|---|---|
| Validated Primary Antibody | ChIP-seq | Immunoprecipitation of target TF. Must be validated for ChIP specificity (e.g., by knockout). |
| Isotype Control IgG | ChIP-seq | Negative control for IP to identify background from antibody Fc region or non-specific bead binding. |
| Protein A/G Magnetic Beads | ChIP-seq | Solid-phase support for antibody-antigen complex capture. Consistency is key for reproducibility. |
| Poly(dI-dC) | EMSA | Non-specific competitor DNA that quenches non-sequence-specific DNA-binding proteins in extracts. |
| γ-³²P ATP or Chemiluminescent Labeling Kit | EMSA | For end-labeling DNA probes to visualize protein-DNA complexes via autoradiography or imaging. |
| Unlabeled Competitor Oligonucleotides | EMSA | Specific and mutant probes for competition assays to define binding sequence specificity. |
| SPRI Beads | ChIP-seq | For consistent post-library purification and size selection, crucial for sequencing quality control. |
| Spike-in Chromatin (e.g., S. pombe, Drosophila) | ChIP-seq | Exogenous chromatin added prior to IP for normalization across samples, correcting for technical variation. |
| RNase A & Proteinase K | Both | Essential enzymes for clean DNA recovery after ChIP or EMSA binding reactions. |
| Non-Denaturing PAGE Gel System | EMSA | Provides the matrix for separation of protein-DNA complexes from free probe based on size/shift. |
ChIP-seq vs EMSA: A Direct Comparison of Strengths, Limitations, and Data Interpretation
Within the broader thesis on ChIP-seq versus EMSA for transcription factor binding research, this in-depth technical guide provides a critical, head-to-head comparison of these two foundational techniques. While EMSA (Electrophoretic Mobility Shift Assay) pioneered the study of protein-nucleic acid interactions in vitro, ChIP-seq (Chromatin Immunoprecipitation followed by sequencing) revolutionized the mapping of these interactions in their native chromatin context in vivo. The choice between them hinges on the specific research question and the trade-offs between throughput, sensitivity, and quantitative capability, which are the central foci of this analysis.
Core Methodologies and Experimental Protocols
EMSA: Detailed Protocol
EMSA, or gel shift assay, detects direct binding of a purified or recombinant transcription factor (TF) to a labeled DNA probe containing a putative binding site.
Key Protocol Steps:
- Probe Preparation: A double-stranded DNA oligonucleotide (typically 20-40 bp) containing the consensus binding sequence is labeled, usually at the 5' end with [γ-³²P]ATP via T4 polynucleotide kinase or with a non-radioactive fluorophore/biotin tag.
- Protein Purification: The TF of interest is expressed and purified from a recombinant system (e.g., E. coli, baculovirus) or sourced from nuclear extracts.
- Binding Reaction: The labeled probe is incubated with the TF protein in a binding buffer (containing MgCl₂, DTT, glycerol, non-specific competitor DNA like poly(dI-dC), and non-ionic detergent) for 20-30 minutes at room temperature.
- Electrophoresis: The reaction mixture is loaded onto a low-ionic-strength, non-denaturing polyacrylamide gel (typically 4-10%). The gel and running buffer (Tris-glycine or Tris-borate) are pre-run and kept cold (4°C).
- Detection & Analysis: The gel is dried and exposed to a phosphorimager screen (radioactive) or directly scanned (fluorescent/chemiluminescent). The shifted protein-DNA complex migrates slower than the free probe.
ChIP-seq: Detailed Protocol
ChIP-seq identifies genome-wide binding sites of a TF or histone modification in living cells.
Key Protocol Steps:
- Crosslinking: Cells are treated with formaldehyde (typically 1%) to covalently crosslink proteins to DNA.
- Cell Lysis & Chromatin Shearing: Cells are lysed, and chromatin is fragmented to 200-600 bp fragments via sonication or enzymatic digestion (MNase).
- Immunoprecipitation (IP): The fragmented chromatin is incubated with a high-specificity antibody against the target TF or histone mark. Antibody-chromatin complexes are isolated using Protein A/G beads.
- Crosslink Reversal & Purification: The immunoprecipitated material is washed, crosslinks are reversed (usually by heating at 65°C), and proteins are digested. The co-precipitated DNA is purified.
- Library Preparation & Sequencing: The DNA fragments are converted into a sequencing library (end-repair, A-tailing, adapter ligation, PCR amplification) and sequenced on a high-throughput platform (e.g., Illumina).
- Bioinformatics Analysis: Reads are aligned to a reference genome. Peak-calling algorithms (e.g., MACS2) identify enriched regions (peaks) compared to a control input sample.
Comparative Analysis: Throughput, Sensitivity, Quantitative Capability
Quantitative Comparison Tables
Table 1: Core Performance Metrics
| Metric | EMSA | ChIP-seq |
|---|---|---|
| Throughput (Sites) | Low (1-10 sites per gel) | Very High (10,000+ genome-wide sites) |
| Sensitivity | High (can detect sub-nanomolar binding affinities in vitro) | Moderate to High (depends on antibody quality, TF abundance, and sequencing depth) |
| Quantitative Capability | Semi-quantitative for affinity (Kd); excellent for kinetics | Semi-quantitative for occupancy; relative enrichment between conditions |
| Context | In vitro, defined sequence | In vivo, native chromatin context |
| Primary Output | Binding confirmation & affinity estimation | Genome-wide binding map & sequence motifs |
| Time to Result | 1-2 days | 3-7 days (wet lab + bioinformatics) |
| Cost per Sample | Low ($100s) | High ($1000s for sequencing) |
Table 2: Technical and Practical Considerations
| Consideration | EMSA | ChIP-seq |
|---|---|---|
| Required Starting Material | Purified protein or nuclear extract | Millions of cells per immunoprecipitation |
| Key Reagent | Labeled DNA probe; purified TF | High-quality, ChIP-validated antibody |
| Artifact Potential | Non-specific shifts; probe purity | Non-specific antibody binding; shearing bias |
| Ability to Detect Cooperative Binding | Yes, with multiple proteins | Indirectly, via motif co-occurrence |
| Dynamic Range | Limited by gel resolution | Several orders of magnitude (via read depth) |
In-Depth Discussion
Throughput: ChIP-seq is the unambiguous winner in throughput, capable of identifying tens of thousands of binding sites across the entire genome in a single experiment. EMSA is inherently low-throughput, designed to interrogate individual, pre-defined DNA sequences.
Sensitivity: Sensitivity definitions differ. EMSA offers exquisite biochemical sensitivity, capable of detecting very low abundance complexes if the binding affinity is high and the probe is hot enough. It can measure dissociation constants (Kd) in the pM-nM range. ChIP-seq's sensitivity is functional; it identifies sites bound in a cellular context but can miss low-affinity or transient binding sites. It is critically dependent on antibody specificity and titer, crosslinking efficiency, and sequencing depth.
Quantitative Capability: Both techniques are primarily semi-quantitative. EMSA can provide quantitative data on relative binding affinities under carefully controlled in vitro conditions using densitometry of gel bands. ChIP-seq data, represented as normalized read counts (e.g., RPKM/FPKM), allows for comparison of relative enrichment across peaks or between samples (e.g., via differential binding analysis tools like DESeq2). However, it does not provide absolute occupancy numbers or direct affinity measurements.
Visualizing Workflows and Context
Diagram 1: ChIP-seq Experimental and Analysis Workflow (78 chars)
Diagram 2: EMSA In Vitro Binding Assay Context (73 chars)
Diagram 3: Decision Logic for Technique Selection (98 chars)
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagent Solutions
| Reagent / Material | Primary Function | Critical Consideration |
|---|---|---|
| ChIP-validated Antibody (ChIP-seq) | Specifically immunoprecipitates the target protein-DNA complex from crosslinked chromatin. | Specificity is paramount. Must be validated for ChIP or ChIP-seq; check vendor citations. |
| Protein A/G Magnetic Beads (ChIP-seq) | Solid support for antibody capture and efficient washing of immunocomplexes. | Consistency in size and binding capacity reduces sample-to-sample variability. |
| Formaldehyde (ChIP-seq) | Reversible crosslinker that fixes protein-DNA and protein-protein interactions in vivo. | Concentration and crosslinking time must be optimized to balance signal and shearing efficiency. |
| Protease/Phosphatase Inhibitors (ChIP-seq) | Preserve the integrity of protein epitopes and complexes during cell lysis and IP. | Essential cocktail to prevent degradation and maintain binding states. |
| Poly(dI-dC) (EMSA) | Non-specific competitor DNA that binds and sequesters non-sequence-specific DNA-binding proteins. | Critical for reducing background; concentration must be titrated for each protein/extract. |
| [γ-³²P]ATP or Chemiluminescent Labeling Kit (EMSA) | Labels the DNA probe for sensitive detection of shifted complexes after electrophoresis. | Radioactivity offers high sensitivity; non-radioactive kits are safer but may be less sensitive. |
| Non-Denaturing Polyacrylamide Gel (EMSA) | Matrix for separating protein-DNA complexes from free probe based on size/charge/shape. | Gel composition (acrylamide:bis ratio) and running conditions (temperature, buffer) are critical. |
| High-Fidelity Taq Polymerase & Seq Adapters (ChIP-seq) | Amplifies and prepares the low-input, purified ChIP DNA for next-generation sequencing. | Reduces PCR bias and ensures efficient adapter ligation for representative library construction. |
This whitepaper examines the concordance and discordance between in vitro and in vivo binding data for transcription factors (TFs), a central challenge in molecular biology and drug discovery. The discussion is framed within the broader methodological comparison of Chromatin Immunoprecipitation followed by sequencing (ChIP-seq) and the Electrophoretic Mobility Shift Assay (EMSA). While ChIP-seq provides a genome-wide, in vivo snapshot of TF occupancy within its native chromatin context, EMSA offers a controlled, in vitro analysis of protein-nucleic acid interactions. Understanding when and why data from these techniques align or diverge is critical for interpreting biological function and validating therapeutic targets.
Core Techniques: EMSA vs. ChIP-seq
Electrophoretic Mobility Shift Assay (EMSA) -In VitroPrinciple
EMSA detects direct binding between a purified or synthesized protein and a labeled DNA or RNA probe via reduced electrophoretic mobility of the complex.
Detailed Protocol:
- Probe Preparation: A short (20-40 bp) double-stranded DNA probe containing the putative binding motif is labeled with a fluorophore or radioisotope (e.g., γ-³²P-ATP).
- Protein Source: Purified recombinant TF or nuclear extract.
- Binding Reaction: The labeled probe is incubated with the protein source in a binding buffer (containing Mg²⁺, KCl, DTT, poly(dI•dC) as non-specific competitor, glycerol) for 20-30 minutes at room temperature.
- Electrophoresis: The reaction mixture is loaded onto a non-denaturing polyacrylamide or agarose gel. The gel and running buffer are pre-chilled (4°C).
- Detection: The gel is imaged using autoradiography, phosphorimaging, or fluorescence. A shifted band indicates a formed complex.
- Controls: Include unlabeled probe in excess (cold competition) to demonstrate specificity and a probe with a mutated binding site.
Chromatin Immunoprecipitation Sequencing (ChIP-seq) -In VivoPrinciple
ChIP-seq identifies genome-wide binding sites of a protein of interest (e.g., a TF) in its native cellular context by crosslinking, immunoprecipitation, and high-throughput sequencing.
Detailed Protocol:
- Crosslinking: Cells are treated with formaldehyde (1%) to covalently link proteins to DNA.
- Cell Lysis & Chromatin Shearing: Cells are lysed, and chromatin is fragmented via sonication to ~200-500 bp fragments.
- Immunoprecipitation: Fragmented chromatin is incubated with an antibody specific to the target TF. Antibody-chromatin complexes are captured using Protein A/G beads.
- Washing & Elution: Beads are washed stringently to remove non-specific binding. Crosslinks are reversed (often at 65°C with high salt), and DNA is purified.
- Library Preparation & Sequencing: Purified DNA fragments are used to generate a sequencing library, which is then sequenced.
- Data Analysis: Sequence reads are aligned to a reference genome. Peak-calling algorithms identify genomic regions with significant enrichment over input control.
Quantitative Data Comparison: Concordance and Discordance
Table 1: Comparative Analysis of EMSA and ChIP-seq
| Aspect | EMSA (In Vitro) | ChIP-seq (In Vivo) | Source of Discordance |
|---|---|---|---|
| Binding Context | Purified components, naked DNA | Native chromatin, nucleosomes, co-factors | Chromatin accessibility & structure |
| Resolution | Single binding site (motif) | 100-500 bp region (peak) | Peak may contain multiple motifs or indirect binding. |
| Throughput | Low (single probe/experiment) | High (genome-wide) | EMSA may miss binding sites discovered by ChIP-seq. |
| Quantitative Output | Binding affinity (Kd), specificity | Enrichment score, peak height | In vivo occupancy influenced by TF concentration and competition. |
| Typical Concordance Rate | ~60-80% for high-affinity canonical motifs within accessible chromatin. | ~20-40% of in vitro motifs may be occupied in vivo due to chromatin constraints. | Varies significantly by TF and cell type. |
Table 2: Factors Causing Discordance Between In Vitro and In Vivo Data
| Factor | Effect on EMSA (In Vitro) | Effect on ChIP-seq (In Vivo) | Outcome |
|---|---|---|---|
| Chromatin Accessibility | Not a factor. | Major determinant; binding only in open/accessible regions. | EMSA predicts binding where chromatin is closed = False Positive. |
| TF Cooperativity | Requires addition of co-factors. | Endogenous co-factors present; cooperative binding common. | EMSA with single TF may show weak/no binding for a cooperative site. |
| Post-Translational Modifications | Often absent in recombinant proteins. | Endogenous PTMs regulate binding affinity & specificity. | Altered binding specificity in vivo vs in vitro. |
| Non-Specific Competition | Simulated with poly(dI•dC). | Complex intracellular milieu of proteins and nucleic acids. | In vitro affinity may not reflect in vivo competitiveness. |
Visualizing the Methodological and Biological Context
Title: Causes of Concordance and Discordance Between EMSA and ChIP-seq Data
Title: Comparative Workflow of EMSA and ChIP-seq Experiments
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for EMSA and ChIP-seq Studies
| Reagent / Kit | Primary Function | Key Consideration for Concordance Studies |
|---|---|---|
| Recombinant Transcription Factor | Purified protein source for EMSA. | Ensure proper folding and presence of critical post-translational modifications if required. |
| Biotin- or Fluorescein-labeled DNA Oligonucleotides | EMSA probe generation. | Labeling method should not interfere with protein-DNA interaction. Cold competitor probes are essential. |
| Poly(dI•dC) | Non-specific competitor DNA in EMSA buffers. | Concentration must be optimized to suppress non-specific binding without outcompeting specific binding. |
| High-Affinity, Validated ChIP-grade Antibody | Immunoprecipitation of target TF in ChIP-seq. | Specificity is paramount; poor antibodies cause high background and false peaks. |
| Chromatin Shearing Reagents (Enzymatic or Sonication) | Fragment chromatin for ChIP. | Fragment size distribution affects resolution; over/under-shearing impacts efficiency. |
| Magnetic Protein A/G Beads | Capture antibody-TF-DNA complexes. | Bead composition affects non-specific binding and wash efficiency. |
| Crosslinking Reversal Buffer | Release DNA from immunoprecipitated complexes. | Complete reversal is necessary for optimal DNA yield and library prep. |
| ChIP-seq DNA Library Prep Kit | Prepare sequencing libraries from low-input, low-complexity DNA. | Kit sensitivity and bias affect detection of lower-affinity binding sites. |
| Spike-in Control DNA/Chromatin | Normalize for technical variation between ChIP samples. | Critical for quantitative comparisons across experiments or conditions. |
Achieving contextual relevance in transcription factor binding research requires a critical, integrated approach. EMSA remains indispensable for defining the intrinsic DNA-binding specificity and affinity of a TF. ChIP-seq reveals the functional, chromatin-contextualized binding landscape within the cell. Discordance is not a failure of either method but a revelation of biological complexity—chromatin barriers, cooperative interactions, and cellular competition. The most robust research strategy employs EMSA to validate high-confidence motifs identified by ChIP-seq and uses ChIP-seq to test the in vivo relevance of in vitro-defined binding sites, thereby closing the loop between biochemical potential and biological reality.
Cost, Time, and Technical Skill Requirements Analysis
Within the critical evaluation of transcription factor (TF) binding research methodologies, the comparative analysis of Chromatin Immunoprecipitation followed by sequencing (ChIP-seq) and Electrophoretic Mobility Shift Assays (EMSA) extends beyond scientific merit to encompass practical constraints. This guide provides a rigorous, technical breakdown of the cost, time, and skill parameters essential for strategic experimental planning in academic and drug discovery settings.
Quantitative Comparative Analysis
Table 1: Direct Cost Breakdown (Approximate USD)
| Component | ChIP-seq (Per Sample) | EMSA (Per Assay) |
|---|---|---|
| Primary Antibody (TF-specific) | $200 - $800 | N/A |
| Protein A/G Magnetic Beads | $30 - $50 | N/A |
| Library Prep Kit | $50 - $150 | N/A |
| Sequencing (5M reads) | $200 - $500 | N/A |
| Radiolabeled (γ-32P) ATP | N/A | $2 - $5 |
| Biotin-labeled Oligo Probe | N/A | $10 - $20 |
| Poly(dI-dC) Competitor DNA | N/A | $1 - $5 |
| Total Estimated Cost | $480 - $1500 | $13 - $30 |
Table 2: Time Investment & Skill Profile
| Parameter | ChIP-seq | EMSA |
|---|---|---|
| Hands-on Time | 3-4 days (discontinuous) | 6-8 hours |
| Total Time to Data | 5-7 days (plus sequencing queue) | 1-2 days |
| Core Technical Skills | Cell culture, chromatin handling, immunoprecipitation, NGS library prep, basic bioinformatics. | Oligo design & annealing, protein extraction/native gel electrophoresis, blotting/detection (chemiluminescent/radioactive). |
| Critical Expertise | Antibody validation, peak-calling analysis, statistical assessment of binding sites. | Optimization of binding conditions, specific vs. non-specific binding differentiation. |
| Automation Potential | Medium (liquid handlers for library prep) | Low (primarily manual) |
Detailed Experimental Protocols
Protocol 1: Key Steps in Native ChIP-seq for TF Binding
- Crosslinking & Harvesting: Treat cells with 1% formaldehyde for 10 min at room temperature. Quench with 125 mM glycine.
- Cell Lysis & Chromatin Shearing: Lyse cells in SDS buffer. Sonicate chromatin to achieve 200-500 bp fragments using a focused ultrasonicator (e.g., Covaris). Confirm size by agarose gel electrophoresis.
- Immunoprecipitation: Pre-clear sheared chromatin with Protein A/G beads. Incubate supernatant with 1-10 µg of validated, TF-specific antibody overnight at 4°C. Capture immune complexes with beads.
- Washing & Elution: Wash beads sequentially with Low Salt, High Salt, LiCl, and TE buffers. Elute complexes in elution buffer (1% SDS, 100 mM NaHCO3).
- Reverse Crosslinks & Purification: Incubate eluate with 200 mM NaCl at 65°C overnight. Treat with RNase A and Proteinase K. Purify DNA using silica membrane columns.
- Library Preparation & Sequencing: Use a commercial kit (e.g., NEBnext) for end-repair, dA-tailing, adapter ligation, and PCR enrichment. Sequence on an Illumina platform (≥5 million reads per sample).
Protocol 2: Key Steps in EMSA with Chemiluminescent Detection
- Probe Preparation: Anneal complementary single-stranded oligonucleotides containing the putative TF binding site. Label the duplex with biotin at the 3' or 5' end using a terminal transferase kit.
- Nuclear Protein Extract Preparation: Harvest cells. Lyse in hypotonic buffer, pellet nuclei. Extract nuclear proteins with high-salt buffer (e.g., 400 mM KCl). Determine protein concentration.
- Binding Reaction: Combine 5-20 µg nuclear extract, 2 µg poly(dI-dC), labeled probe (20 fmol), in binding buffer (10 mM Tris, 50 mM KCl, 1 mM DTT, 10% glycerol). Incubate 20-30 min at room temperature.
- Gel Electrophoresis: Load reaction onto a pre-run 6% non-denaturing polyacrylamide gel in 0.5X TBE buffer. Run at 100V for 60-90 min at 4°C.
- Transfer & Detection: Electroblot DNA-protein complexes to a positively charged nylon membrane. Crosslink DNA to membrane via UV. Detect using streptavidin-HRP and chemiluminescent substrate, imaging with a CCD camera.
Visualization of Methodological Workflows
Title: ChIP-seq Experimental Workflow
Title: EMSA Experimental Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for TF Binding Studies
| Item | Function & Relevance |
|---|---|
| Validated ChIP-grade Antibody | Crucial for ChIP-seq specificity. Must be validated for immunoprecipitation under cross-linked conditions. Target: Transcription factor of interest. |
| Magnetic Protein A/G Beads | Enable efficient capture and washing of antibody-chromatin complexes in ChIP-seq, reducing non-specific background. |
| Chromatin Shearing Reagents (Covaris microTUBEs) | Optimized for acoustic shearing to yield consistent fragment sizes (200-500 bp) for high-resolution ChIP-seq. |
| NGS Library Preparation Kit | Streamlines conversion of immunoprecipitated DNA into sequencer-compatible libraries (e.g., Illumina TruSeq). |
| Biotin 3' End DNA Labeling Kit | Provides a non-radioactive, stable method to label EMSA probes for chemiluminescent detection. |
| Poly(dI-dC) Competitor DNA | Critical for EMSA to suppress non-specific protein-DNA interactions, enhancing specificity of the shifted band. |
| Non-denaturing Polyacrylamide Gel Mix | Forms the matrix for EMSA separation based on protein-DNA complex size/charge. |
| Chemiluminescent Nucleic Acid Detection Module | For sensitive, non-radioactive visualization of biotin-labeled EMSA probes after transfer. |
The choice between ChIP-seq and EMSA is dictated by the research question's scope balanced against resource constraints. ChIP-seq delivers genome-wide, in vivo binding profiles but demands significant investment in cost, time, and computational expertise. EMSA offers an economical, rapid, and accessible in vitro validation tool for focused, mechanistic studies of specific protein-DNA interactions but lacks genomic context. A synergistic approach, using EMSA to validate ChIP-seq-predicted binding sites, often represents the most robust strategy within a comprehensive thesis on TF binding research.
In transcription factor (TF) binding research, the integration of high-throughput discovery (ChIP-seq) and low-throughput validation (EMSA) is fundamental for establishing rigorous, reproducible results. This guide details the complementary validation strategies, providing protocols, data interpretation frameworks, and practical toolkits for researchers and drug development professionals.
The broader thesis posits that while Chromatin Immunoprecipitation followed by sequencing (ChIP-seq) is the de facto standard for genome-wide mapping of TF binding events, its results are inherently probabilistic and can include false positives due to antibody cross-reactivity or bioinformatic peak-calling artifacts. Conversely, the Electrophoretic Mobility Shift Assay (EMSA) provides direct, in vitro biochemical evidence of protein-DNA interaction but lacks genomic context and cellular complexity. Therefore, a cyclical validation strategy—using EMSA to confirm specific ChIP-seq-identified sequences and using prior EMSA-validated motifs to inform ChIP-seq analysis—is critical for robust scientific conclusions.
Core Principles of Each Technique
ChIP-seq: Genome-Wide Discovery
ChIP-seq crosslinks proteins to DNA in vivo, immunoprecipitates the protein of interest with its bound DNA fragments, and sequences them. Peaks represent enriched genomic regions.
Key Quantitative Outputs:
- Peak Score: -log10(p-value) or -log10(q-value) from peak callers (MACS2, HOMER).
- Fold Enrichment: Signal over control (IgG or Input).
- Peak Location: Promoter, enhancer, intron, etc.
EMSA:In VitroBiochemical Validation
EMSA detects direct binding of a purified or in vitro transcribed/translated protein to a labeled DNA probe via a gel shift in electrophoretic mobility.
Key Quantitative Outputs:
- Dissociation Constant (Kd): Measures binding affinity.
- Shift Intensity: Percentage of probe shifted, indicating binding efficiency.
Quantitative Data Comparison
Table 1: Comparative Analysis of ChIP-seq and EMSA
| Parameter | ChIP-seq | EMSA |
|---|---|---|
| Throughput | High-throughput (genome-wide) | Low-throughput (single sequence/probe) |
| Binding Context | In vivo (cellular environment, chromatin context) | In vitro (purified components, no chromatin) |
| Primary Output | Genomic peak coordinates, motif enrichment | Binary yes/no for binding, affinity measurement (Kd) |
| Key Strength | Unbiased discovery, genomic localization, allelic-specific binding detection | Direct binding confirmation, affinity/ specificity quantification, mutant analysis |
| Key Limitation | Indirect (relies on antibody), false positives/negatives possible | Does not confirm in vivo binding, may miss co-factor requirements |
| Typical Timeline | 1-2 weeks (from cells to data) | 1-3 days |
| Approximate Cost per Sample | High (sequencing costs) | Low |
Table 2: Validation Metrics from Integrated Studies (Hypothetical Data Summary)
| ChIP-seq Peak Set | Peaks Tested by EMSA | EMSA-Validated Peaks | Validation Rate | Common Kd Range (EMSA) |
|---|---|---|---|---|
| High-confidence (q<0.01) | 20 | 18 | 90% | 0.1 - 5 nM |
| Low-confidence (q<0.05) | 20 | 8 | 40% | 5 - 50 nM |
| Negative Genomic Regions | 10 | 1 | 10% | >100 nM |
Detailed Experimental Protocols
Protocol: Validating ChIP-seq Peaks with EMSA
A. Probe Design from ChIP-seq Data:
- Select Peaks: Choose top-ranked peaks (by p-value) and include negative control regions (non-peak, gene desert).
- Extract Sequences: Isolate ±50 bp around the peak summit (from FASTA file).
- Design Oligos: Design complementary 20-40 bp oligonucleotides containing the putative motif. Add 5-10 bp flanking sequences. Include a 5' overhang for labeling (e.g., GATC overhang for Klenow fill-in).
B. EMSA Procedure:
- Probe Labeling:
- Anneal oligos.
- Label with [γ-³²P]ATP using T4 Polynucleotide Kinase or via fill-in reaction with [α-³²P]dCTP and Klenow fragment.
- Purify labeled probe using a spin column (e.g., G-25 Sephadex).
- Protein Preparation: Use purified recombinant TF protein or nuclear extract.
- Binding Reaction (20 µL):
- Labeled Probe: 10,000-20,000 cpm (~0.1-1 ng)
- Protein: 0.5-10 µg nuclear extract or 10-200 ng recombinant protein
- Poly(dI·dC): 1-2 µg (non-specific competitor)
- Binding Buffer: 10 mM HEPES (pH 7.9), 50 mM KCl, 1 mM DTT, 0.5 mM EDTA, 10% glycerol.
- Incubate at 25°C for 20-30 minutes.
- Competition Assay (Specificity Control):
- Include 50-200x molar excess of unlabeled wild-type (specific) or mutated (non-specific) competitor DNA.
- Supershift Assay (Identity Control):
- Add 1-2 µg of specific antibody to the binding reaction. A further shift ("supershift") confirms TF identity.
- Electrophoresis:
- Load reaction onto a pre-run 4-6% non-denaturing polyacrylamide gel in 0.5x TBE buffer.
- Run at 100-150 V at 4°C for 1.5-2 hours.
- Dry gel and expose to phosphorimager screen or X-ray film.
Protocol: Informing ChIP-seq with Prior EMSA Data
- Motif Construction: Use sequences validated by EMSA to create or refine a Position Weight Matrix (PWM) via tools like MEME.
- Peak Filtering/Scoring: Use this PWM in ChIP-seq analysis pipelines (e.g., HOMER
findMotifsGenome.pl, MEME-ChIP) to:- Filter peaks lacking the validated motif.
- Score and rank peaks based on motif match strength.
- Validation Loop: Select new peaks containing the strong motif matches for subsequent EMSA validation.
Visualization of Workflows and Relationships
Title: Cyclical Workflow for ChIP-seq and EMSA Integration
Title: Detailed EMSA Experimental Workflow for Validation
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Integrated ChIP-seq/EMSA Studies
| Item / Reagent | Function / Purpose | Example Product / Note |
|---|---|---|
| ChIP-Grade Antibody | Specific immunoprecipitation of the target TF in its native chromatin context. Critical for ChIP-seq specificity. | Validated antibodies from Abcam, Cell Signaling, Diagenode. |
| Proteinase K | Digests proteins post-crosslinking reversal in ChIP. Essential for clean DNA recovery. | Molecular biology grade, RNA-free. |
| Magnetic Protein A/G Beads | Efficient capture of antibody-TF-DNA complexes for ChIP. Enable easy washes. | Dynabeads (Thermo Fisher). |
| High-Fidelity DNA Polymerase | Amplifies ChIP-enriched DNA for library preparation. Minimizes PCR bias. | KAPA HiFi HotStart ReadyMix. |
| [γ-³²P]ATP or Biotin-labeled dNTPs | Radiolabels or tags EMSA DNA probes for sensitive detection. | PerkinElmer; Biotin labeling kits (Thermo Fisher). |
| Recombinant TF Protein | Provides a pure, consistent protein source for EMSA binding reactions and affinity measurements. | Expressed and purified in-house or purchased (e.g., Active Motif). |
| Poly(dI·dC) | Non-specific competitor DNA in EMSA. Reduces non-specific protein-probe interactions. | Salmon sperm DNA is an alternative. |
| Non-denaturing PAGE Gel System | Matrix for separating protein-DNA complexes from free probe based on size/charge shift in EMSA. | Mini-PROTEAN Tetra Cell (Bio-Rad). |
| Phosphorimager Screen & Scanner | Highly sensitive detection and quantification of radiolabeled EMSA results. | Typhoon FLA systems (Cytiva). |
| MEME Suite / HOMER Software | Discovers de novo motifs (MEME) and performs comprehensive ChIP-seq peak analysis & motif finding (HOMER). | Open-source bioinformatics tools. |
Within the framework of evaluating ChIP-seq versus EMSA for transcription factor (TF) binding research, the selection of a target identification method is a pivotal first step in drug discovery. This whitepaper presents technical case studies demonstrating how complementary techniques are applied to elucidate drug targets and mechanisms of action, directly impacting the development of novel therapeutics.
Core Methodologies in Target Identification
Chromatin Immunoprecipitation Sequencing (ChIP-seq)
Protocol: Cells are cross-linked with formaldehyde, chromatin is sheared via sonication, and TF-bound DNA fragments are immunoprecipitated using a target-specific antibody. After reverse cross-linking, the purified DNA is used to construct a sequencing library. High-throughput sequencing and peak-calling algorithms identify genomic binding sites.
Case Study: Targeting BET Bromodomains in Oncology
- Objective: Identify transcriptional programs regulated by BRD4 to understand the mechanism of BET inhibitors like JQ1.
- Application: ChIP-seq for BRD4 and histone marks (H3K27ac) in leukemic cells before/after JQ1 treatment.
- Data Outcome: Revealed specific displacement of BRD4 from super-enhancers of key oncogenes (e.g., MYC), providing a mechanistic rationale for therapeutic efficacy.
Electrophoretic Mobility Shift Assay (EMSA)
Protocol: A purified protein or nuclear extract is incubated with a labeled DNA probe containing a putative binding sequence. The reaction mixture is loaded onto a non-denaturing polyacrylamide gel. Protein-DNA complexes exhibit reduced electrophoretic mobility ("shift") compared to free probe, confirmed via competition with unlabeled probe or supershift with an antibody.
Case Study: Validating NF-κB Inhibitor Mechanisms
- Objective: Confirm direct inhibition of NF-κB p65 subunit DNA binding by a small molecule candidate.
- Application: EMSA using recombinant p65 and a consensus κB site probe, with/without the inhibitor.
- Data Outcome: Quantitative reduction in band shift intensity demonstrated direct, dose-dependent disruption of DNA binding, a key mechanistic validation step.
Affinity-Based Proteomics (Chemoproteomics)
Protocol: A drug molecule is immobilized on a solid support to create a bait. Incubation with cell lysates allows binding of interacting proteins, which are then eluted, digested, and identified by mass spectrometry (LC-MS/MS).
Case Study: De-orphaning a Phenotypic Hit
- Objective: Identify the cellular target of a compound inducing apoptosis in cancer cells.
- Application: Use of the compound tethered to beads for pull-down, followed by quantitative MS versus control beads.
- Data Outcome: Identification of a previously uncharacterized interaction with a mitochondrial enzyme, revealing a novel apoptotic pathway.
CRISPR-Cas9 Screening
Protocol: A genome-wide library of guide RNA (gRNA)-expressing lentiviruses is used to generate a pool of knockout cells. This population is subjected to a selective pressure (e.g., drug treatment). Deep sequencing of gRNAs pre- and post-selection reveals enriched or depleted guides, indicating genes whose loss confers resistance or sensitivity.
Case Study: Identifying Synthetic Lethal Partners for KRAS Mutants
- Objective: Find genes essential for the survival of KRAS-mutant but not wild-type colorectal cancer cells.
- Application: Parallel genome-wide CRISPR knockout screens in isogenic KRAS mutant vs. WT cell lines.
- Data Outcome: Identification of several kinases whose knockout selectively inhibited mutant cell proliferation, nominating new co-targets.
Table 1: Quantitative Comparison of Core Target ID Methods
| Method | Primary Output | Throughput | Sensitivity | Key Quantitative Metric | Typical Timeline |
|---|---|---|---|---|---|
| ChIP-seq | Genome-wide binding loci | High | High (needs antibody) | Peak enrichment FDR, read counts | 5-7 days |
| EMSA | Confirmation of direct binding in vitro | Low | Moderate | Band shift intensity (densitometry) | 1-2 days |
| Chemoproteomics | Direct protein interactors | Medium | High (depends on bait) | Spectral counts, fold-enrichment | 1-2 weeks |
| CRISPR Screen | Genes affecting phenotype | Very High | High | gRNA fold-change, MAGeCK score | 3-4 weeks |
Integrated Mechanistic Study Workflow
Diagram 1: Integrated drug discovery workflow from target ID to mechanism.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Featured Experiments
| Reagent / Kit | Primary Function | Example Use Case |
|---|---|---|
| Magna ChIP Kit | Optimized buffers & beads for chromatin IP. | ChIP-seq sample preparation for histone marks/TFs. |
| LightShift Chemiluminescent EMSA Kit | Provides biotin-labeling, binding, and detection reagents. | Validating TF-inhibitor interactions via non-radioactive EMSA. |
| CellTiter-Glo Luminescent Viability Assay | Measures ATP as a proxy for metabolically active cells. | Assessing cell viability post-treatment in phenotypic screens. |
| Pierce Magnetic Agarose for Pull-Down | Beads for immobilizing bait molecules. | Chemoproteomics target identification studies. |
| LentiCRISPR v2 Plasmid | All-in-one lentiviral vector for gRNA & Cas9 expression. | Construction of libraries for CRISPR-Cas9 knockout screens. |
| NE-PER Nuclear Extract Kit | Fractionates cell lysates to isolate nuclear proteins. | Provides protein extract for EMSA or TF activity assays. |
| TruSeq ChIP Library Prep Kit | Prepares immunoprecipitated DNA for sequencing. | Generating sequencing libraries from ChIP DNA fragments. |
Comparative Analysis: ChIP-seq vs. EMSA in TF Research
The core thesis distinguishing ChIP-seq and EMSA lies in their application scope: discovery versus reductionist validation.
- ChIP-seq provides an unbiased, genome-wide view of TF occupancy in its native chromatin context, including co-factor interactions. It is indispensable for identifying novel regulatory regions and understanding epigenetic mechanisms in disease states.
- EMSA offers a highly specific, biophysical assay to confirm direct, sequence-specific binding of a purified TF. It is optimal for dissecting binding kinetics, assessing mutant protein function, or screening for direct DNA-binding inhibitors.
Diagram 2: Decision flow for ChIP-seq vs EMSA based on research question.
Table 3: Strategic Application in Drug Discovery
| Stage | Preferred Method (ChIP-seq vs EMSA) | Rationale |
|---|---|---|
| Target Identification | ChIP-seq (or CRISPR screen) | Unbiased discovery of oncogenic TF hubs and cis-regulatory networks. |
| Mechanism of Action | ChIP-seq (pre/post treatment) | Maps global changes in TF occupancy and histone modifications upon drug treatment. |
| Hit-to-Lead Optimization | EMSA | High-throughput capability to rank analogs by direct target engagement potency. |
| Preclinical Biomarker | ChIP-seq (on patient samples) | Defines pathogenic enhancer signatures predictive of drug response. |
Effective drug discovery requires the strategic application of complementary techniques. While CRISPR screens and chemoproteomics excel at initial target deconvolution, and EMSA provides crucial reductionist validation of direct binding, ChIP-seq stands apart for elucidating the in vivo transcriptional mechanisms of both disease drivers and therapeutic interventions. The choice between ChIP-seq and EMSA is not one of superiority but is defined by the specific biological question—be it genome-wide discovery or precise biochemical validation—within the mechanistic pipeline.
Conclusion
ChIP-seq and EMSA are not mutually exclusive but rather complementary pillars in the study of transcription factor biology. EMSA remains the gold standard for definitive, quantitative in vitro validation of specific protein-DNA interactions, offering mechanistic insight through mutagenesis. ChIP-seq provides the indispensable genome-wide, in vivo context, revealing the full landscape of TF occupancy and its correlation with gene expression. For robust conclusions, especially in translational research and drug development, a synergistic approach is recommended: using ChIP-seq for unbiased discovery and EMSA for focused, mechanistic validation of key targets. Future directions, including the integration of CUT&Tag for low-input samples and advanced computational models, will further refine our ability to decode transcriptional regulation, accelerating the development of novel therapeutics targeting dysregulated transcription factors in disease.